TLDR: fastai2 is great, but if you use numpy arrays as input it may be slower than fastai v1. However, if you use NumpyDatasets/ NumpyDataLoader your training will be 33% faster than v1 and 40+% faster than v2.
During the last couple of weeks, I’ve been porting timeseriesAI to fastai v2.
timeseriesAI is a Practical Deep Learning for Time Series / Sequential Data package I built on top fastai v1, that takes X (and y) numpy arrays as input.
The new package can be easily installed from pip (`pip install tsai).
I’ve learned a few things in this process I’d like to share with you.
I think fastai v2 is great. It has many advantages over v1 (like batch tfms, fine-tuning, callbacks, etc). I also think the layered approach allows us to adapt the library to our needs. I used the vanilla datablock API and it worked fine. It’s easy to use and very flexible. 
The fastai ecosystem is superb! nbdev is super useful: a great tool to develop, test and distribute code. Thanks to nbdev I’ve built my first ever pip package. I couldn’t believe how easy it was.
And fastcore has some very useful code too.
Thanks a lot Jeremy and Sylvain for putting all this together!) 


When I learned how to use fastai v2, I run some performance tests with time series data (numpy arrays) to compare v1 and v2. I was disappointed by the poor results:
- v2 training is 40-60% slower.
- v2 dataloader takes 2-3x to return batches.
- v2 dataset takes 2x to return items.
fastai v2 is excellent when using images, text, tabular data, etc, but it’s not streamlined to use numpy arrays. 
Since I use fastai on a daily basis, I started to investigate if there was any way to speed up the code for numpy arrays.
I found a couple of modifications that could potentially accelerate batch creation (1) as long as:
- No item tfms are applied to X (and y), or
- Item tfms are deterministic (non-random) and its output fits in memory
These conditions are present in many time series problems.
These 2 processes are:
- Apply item tfms inplace (dataset): in time series problems, most item transforms are deterministic (non-random), which means they can be applied during dataset initialization, instead of at batch creation time. In this way they are only applied once instead of once per epoch.
- Get all batch items at once instead of one by one (dataloader): if you carefully build the output during datasets initialization, you only need to slice X (and y) and cast them to the desired output types (tensor, TensorCategory, etc.) to create a batch. Slicing and casting are very fast. This removes the need to have a collate function. The only thing the dataloader needs to do is to pass the indices that will be applied in each iteration to get a batch.
With these modifications, the batch creation process with numpy arrays is super fast: 100 times faster than vanilla v2, and 30 times faster than v1. 
The user interface I built is very similar to scikit-learn’s API:
NumpyDatasets(X, y=None, tfms=tfms, splits=splits, inplace=True)
TSDatasets(X, y=None, tfms=tfms, sel_vars=None, sel_steps=None, splits=splits, inplace=True)
I’ve created a couple of notebooks to the timeseriesAI repo to show you how all this works.
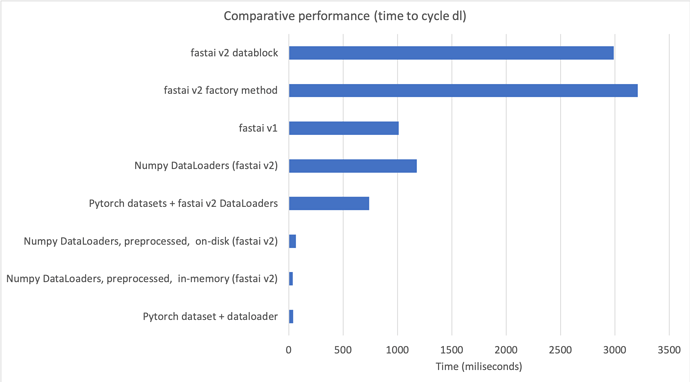
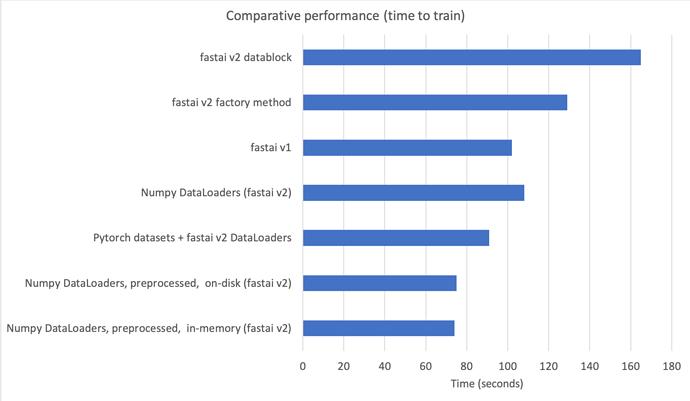
In summary, this means we can have all the benefits of v2 when using numpy arrays with a simple scikit-learn-like API, that is 43-55% faster than factory methods and datablock API in v2, and 30% faster than v1 (see 00b_How_to_use_numpy_arrays_in_fastai2.ipynb)
This is the comparison in a chart:
If you are interested I’d suggest you start using this introductory nb.
If you decide to try it, it’s be good if you would provide some feecback on how it works.
 thanks for sharing!
thanks for sharing!