Yes, that’s better. 007b takes in itself 4-5 hours to run on a p3!
Pushed a few commits here and there to refactor a lot of the NLP stuff.
The idea is to have the data loaded and a learner in just a few lines of code, like in CV.
Merged docstrings branch and just added another PR here.=
Preview of core.py - (this example will not be checked in)
Summary:
Reformatted function/class/enum definition.
Trying to provide links where possible - inside docstrings, subclasses
Show global variables in documentation notebooks FileLike = Union[str, Path]
Next:work on making sure links go to correct places and formatting the html
Fixed a bug in yesterday’s implementation of separating batchnorm layers for weight decay in this commit.
There is now a flag bn_wd in Learner which, if set to False, will prevent weight decay from being applied to batchnorm layers during training.
In this commit, created an ImageBBox object to get data augmentation working with bounding boxes.
Hunder the hood, it’s just a square mask and when we need to pull the data at the end, we take the min/max of the coordinates of non-zero elements.
Got a bit behind on updates. Here we go:
- Add titles to show_image
-
Split a
Collectionrandomly - Add test sets
-
data_from_imagefoldercreates a DataBunch for you from a folder of folders of folders! - Changed transform defaults to be more useful
- TTA
-
Only include image extensions in
get_image_files - You can override this with
check_ext=False - Various metrics and loss functions now flatten their params
- Figure out how many CPUs to use
- Added a model meta dictionary so that models know how to cut/split themselves
In 002_images.ipynb there is a very complex chain of ands, ors and nots (#1):
def get_image_files(c:Path, check_ext:bool=True)->FilePathList:
[...]
return [o for o in list(c.iterdir())
if not o.name.startswith('.') and not o.is_dir()
and (not check_ext or (o.suffix in image_extensions))]
I had a bit of a smoke coming up parsing the last line in my head.
Won’t this be more readable (#2):
if not o.name.startswith('.') and not o.is_dir()
and not (check_ext and o.suffix not in image_extensions)
And then it allows us to drop 2 nots (#3), but the above is fine too - it’s consistent on negating everything and there are less parenthesis:
if not (o.name.startswith('.') or o.is_dir()
or (check_ext and o.suffix not in image_extensions))
Too bad python doesn’t have unless 
It’s type-annotation Friday!
- annotated nb 009 in this commit
- annotated nb 008 in this commit
- annotated the 007 nbs in this commit
- made a few corrections to early notebooks in this commit
Continue to clean-up with
- moving 009a to x_009
- solving a bug with flat_master and the latest no wd to bn layers development in 004b
Yup that looks better to me.
Also @313V has been adding type annotations and docstrings to the earlier notebooks.
Didn’t have time to post a message here yesterday, but the modules have been added in this commit
I made a few changes this morning in this commit then corrected bugs and added the all_ for each module that needs it in this commit.
Finally in this commit I added five examples notebooks to check everything was working well (dogs and cats, cifar10, imdb classification, movie lens and rossmann).
As Jeremy explained, you shouldn’t touch the dev_nb anymore (except to add prose). Bug fixes should be done in the modules directly! You should also use a pip install -e of the new library to test those notebooks, to easily have the latest version installed.
One last commit about module developments for a while. Just added mixup that allows us to get very fast results on cifar10 (6 minutes for 94% accuracy).
After the recent rewrite of history you may get this error when running: git pull (or direct merge)
fatal: refusing to merge unrelated histories
The easiest way to fix this if you have forked fastai/fastai_v1 and you don’t have any branches that you want to keep, is to nuke your fork by following the delete option at the end of:
https://github.com/<yourusername>/fastai_v1/settings
and then forking again.
A potentially much more complex way is to (assuming you use ssh, adjust for https: urls if need be):
git clone git@github.com:YOURUSER/fastai_v1.git
cd fastai_v1
git remote add upstream git@github.com:fastai/fastai_v1.git
git fetch upstream
git checkout master
git merge upstream/master --allow-unrelated-histories
which depending on when you synced your forked repository with fastai/fastai_v1 may create a gazillion of conflicts or not. In my case it did, so I decided re-doing the fork is the easiest option.
Once you resolved the conflicts, push back in to sync it:
git push --set-upstream origin master
If you don’t use the forked project, but a direct fastai/fastai_v1 checkout you can use -allow-unrelated-histories with git pull, or simply make a new checkout and copy your work files over to the fresh repo checkout.
The initial documentation website commit has been done - we’ve simply imported the standard jekyll documentation template at this point, and are now working on filling it in with our docs. So everything inside doc/ now is from the template. Once we’ve figured out what we need we can remove some of the redundant stuff.
I originally checked in the vendor/ directory for jekyll, but changed my mind after feedback from @stas, which is why I had to rewrite history - see the previous message from @stas if that causes any problems for you (should only have a problem if you have a fork).
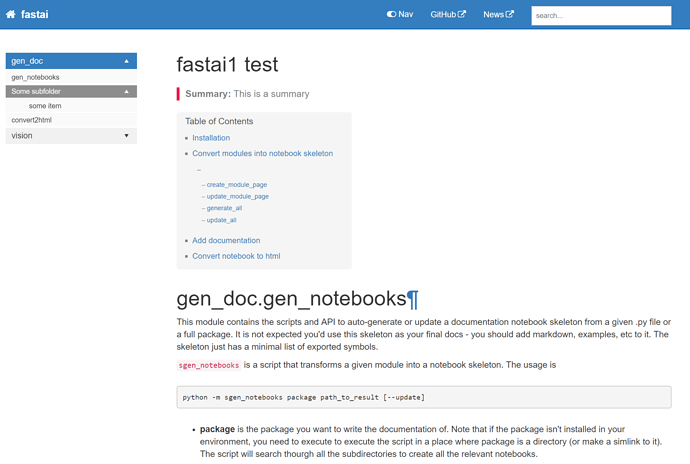
Yay it looks like this doc template may just work nicely! I just manually popped in one page for testing, and it’s looking pretty good without even customizing anything much at all:
Hmm, are you able to run dev_nb/002_images.ipynb?
I have to change the first cell to even find gen_doc:
- import sys
- sys.path.append('../docs')
to:
+ import pathlib, sys
+ path = str((pathlib.Path(".")/".."/"fastai").resolve())
+ if path not in sys.path: sys.path.insert(0, path)
there are no python libs under fastai_v1/docs, not sure how it worked…
and then once the path has been fixed it fails internally:
---------------------------------------------------------------------------
ValueError Traceback (most recent call last)
<ipython-input-2-2e0a20ad4bf5> in <module>()
7 if path not in sys.path: sys.path.insert(0, str(path))
8 sys.path
----> 9 from gen_doc.nbdoc import show_doc as sd
/mnt/disc1/fast.ai-1/br/fastai/master/fastai/gen_doc/nbdoc.py in <module>()
3 from typing import Dict, Any, AnyStr, List, Sequence, TypeVar, Tuple, Optional, Union
4 from .docstrings import *
----> 5 from .core import *
6
7 __all__ = ['get_class_toc', 'get_fn_link', 'get_module_toc', 'show_doc', 'show_doc_from_name',
/mnt/disc1/fast.ai-1/br/fastai/master/fastai/gen_doc/core.py in <module>()
----> 1 from ..core import *
2 import re
3
4 def strip_fastai(s): return re.sub(r'^fastai\.', '', s)
5
ValueError: attempted relative import beyond top-level packageMade some little moves in this commit
- It made more sense to have DatasetBase and LabelDataset in data
- I renamed all the data_from_* function to something more consistent like {type}_data_from_* so for instance image_data_from_folder, text_data_from_tokens or tabular_data_from_df.
- Then I changed all the references to those functions in the example notebooks.
Fixed now.
Hmm, are you able to run
dev_nb/002_images.ipynb
Fixed now.
ModuleNotFoundError: No module named 'fastai'
Are notebooks now supposed to rely on a pre-installed fastai as being discussed in the other thread?
Otherwise the following would remove such requirement:
import pathlib, sys
path = str((pathlib.Path(".")/"..").resolve())
if path not in sys.path: sys.path.insert(0, path)