Can someone expline me what every line in the loop does:
print(ss,"\n")
for i in range(50):
n=res[-1].topk(2)[1]
n = n[1] if n.data[0]==0 else n[0]
print(TEXT.vocab.itos[n.data[0]], end=' ')
res,*_ = m(n[0].unsqueeze(0))
print('...')
When I run it I get the following error:
/home/paperspace/anaconda3/envs/fastai/lib/python3.6/site-packages/ipykernel_launcher.py:5: UserWarning: invalid index of a 0-dim tensor. This will be an error in PyTorch 0.5. Use tensor.item() to convert a 0-dim tensor to a Python number
"""
---------------------------------------------------------------------------
ValueError Traceback (most recent call last)
<ipython-input-34-d2cf2df24465> in <module>()
4 n = n[1] if n.data[0]==0 else n[0]
5 print(TEXT.vocab.itos[n.data[0]], end=' ')
----> 6 res,*_ = m(n.unsqueeze(0))
7 print('...')
~/anaconda3/envs/fastai/lib/python3.6/site-packages/torch/nn/modules/module.py in __call__(self, *input, **kwargs)
489 result = self._slow_forward(*input, **kwargs)
490 else:
--> 491 result = self.forward(*input, **kwargs)
492 for hook in self._forward_hooks.values():
493 hook_result = hook(self, input, result)
~/anaconda3/envs/fastai/lib/python3.6/site-packages/torch/nn/modules/container.py in forward(self, input)
89 def forward(self, input):
90 for module in self._modules.values():
---> 91 input = module(input)
92 return input
93
~/anaconda3/envs/fastai/lib/python3.6/site-packages/torch/nn/modules/module.py in __call__(self, *input, **kwargs)
489 result = self._slow_forward(*input, **kwargs)
490 else:
--> 491 result = self.forward(*input, **kwargs)
492 for hook in self._forward_hooks.values():
493 hook_result = hook(self, input, result)
~/fastai/courses/dl1/fastai/lm_rnn.py in forward(self, input)
91 dropouth, list of tensors evaluated from each RNN layer using dropouth,
92 """
---> 93 sl,bs = input.size()
94 if bs!=self.bs:
95 self.bs=bs
ValueError: not enough values to unpack (expected 2, got 1)
And I do not really understand the code, therefore I don’t know how to fix it.
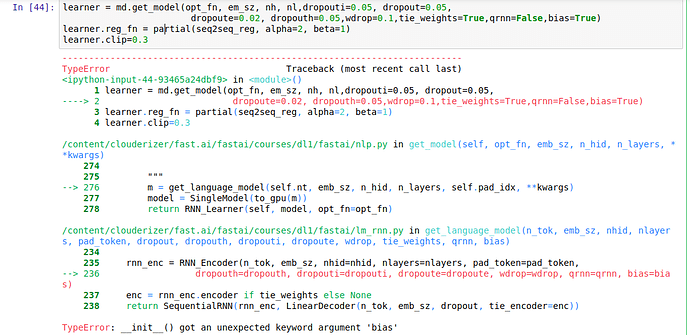
Another problem i have is when running the following code:
m=learner.model
ss=""". So, it wasn't quite was I was expecting, but I really liked it anyway! The best"""
s = [spacy_tok(ss)]
t=TEXT.numericalize(s)
' '.join(s[0])
I get:
TypeError: sequence item 0: expected str instance, spacy.tokens.token.Token found
I believe