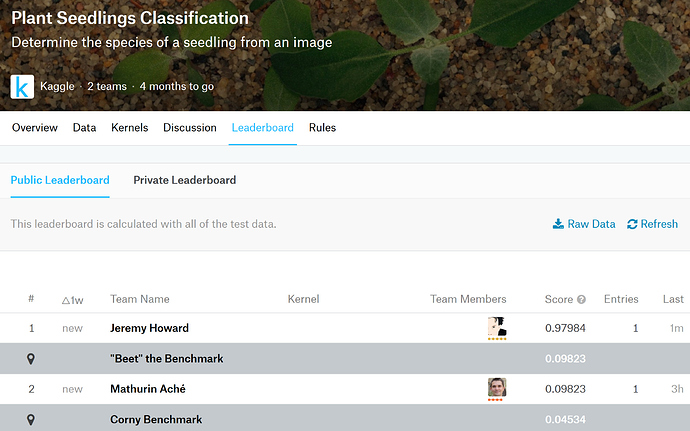
This one looks beginner friendly, yet the images are sufficiently different than the ones in ImageNet to make it interesting.
@jeremy
Looks like this is the metric that I need to include in the evaluation.
from sklearn.metrics import f1_score
f1_score(data.val_y, preds, average=‘micro’)
What is the way to add it to the model??
@gerardo please don’t at-mention me unless you specifically need me to answer your question and no-one else. There’s lots of folks here who can be helpful! 
Look at how ‘metrics’ is defined in the various lesson notebooks we’ve seen so far to see how we’ve done this - especially the Planet one.
I’m sorry to bother. 
I just feel this weird attachment to your work and I have you “cyber-close” that I have the urge to ask questions.
I’m pretty sure that I’m not alone.
I will keep asking questions while I’m keep looking for the them in the forums.

Thanks for all the hard work.
No problem!
Hey i know its silly…
But how do you divide the train folder into valid (i mean getting everything perfectly into sub-folders).
I did it manually
200 images per category for training
the rest for validation.
@gerardo
hmmm I would love to know if there’s any shortcut script if someone wrote it…
else I know what to do next 
I created a CSV using python and then used from_csv.
I also wrote a script to create a labels.csv file with headers file,species.
from glob2 import glob
import pandas as pd
df = pd.DataFrame(columns=["file", "species"])
for image in glob("train/**/*.png"):
dir_ = image.split('/')
file_, species = dir_[-1], dir_[-2]
df = df.append({
"file": file_,
"species": species
}, ignore_index=True)
df.to_csv('labels.csv', index=False)
Then, you can use the from_csv method.
Once you are done creating labels.csv, don’t forget to remove the species folders in train. Keep the images, remove the folders.
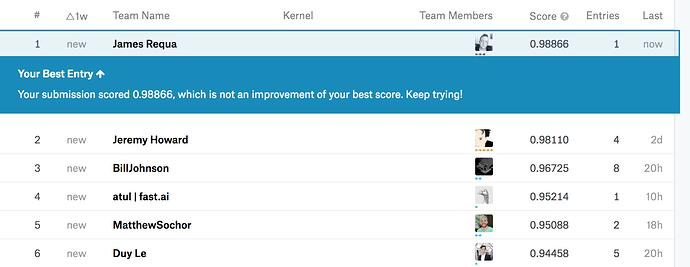
Wow you killed it! What architecture are you using, if I may ask?
Resnet50…how about you?
Also resnet50. Oddly enough I found Resnext50 and Nasnet both a little worse (Resnext50 was better on validation, but worse when submitted).
I tried resnext50 first cause I also thought it would do better, but kept getting cuda out of memory errors on fine-tuning so thats how I ended up resorting to resnet50…will test out a few others. Also still need to train rn50 with all the data or cv, this is just one fold for now
Wow that’s impressive - mine is trained with all data.
To train the bigger models I decreased the batch size. For Nasnet I need bs=12 on a 16GB P3!
Hey how did you include metrics in?
This is what i tried to do.
from sklearn.metrics import f1_score
def meanfscore(y_pred, targ):
return f1_score(targ, y_pred, average='micro')
learn.fit(0.01, 5, metrics=[meanfscore])
This should work but it’s throwing a weird error.
ValueError: Classification metrics can't handle a mix of multilabel-indicator and continuous-multioutput targets
Personally, I didn’t use f1_score as a metric during training. I just used regular accuracy. The LB is scored by f1 based on the predictions you submit.
log_preds,y = learn.TTA()
preds = np.exp(log_preds)
preds = np.argmax(log_preds, axis=1)
metrics.f1_score(data.val_y, preds, average=‘micro’)
I used this code to calculate the f1_score but I would like to add it the learn.fit but does not seems to be working.