About my data:
205205 rows × 495 columns
Task: binary classification.
Data setup:
data = TabularDataBunch.from_df(".", df, dep_var="target", bs=1000,
cont_names=cont_vars, cat_names=cat_vars,
valid_idx=valid_idx, procs=[FillMissing, Categorify, Normalize])
Learner setup:
emb_sizes = {x:10 for x in cat_vars}
learn = tabular_learner(data, layers=[200, 100], emb_drop=0.04, emb_szs=emb_sizes,
ps=[0.001,0.01], metrics=accuracy)
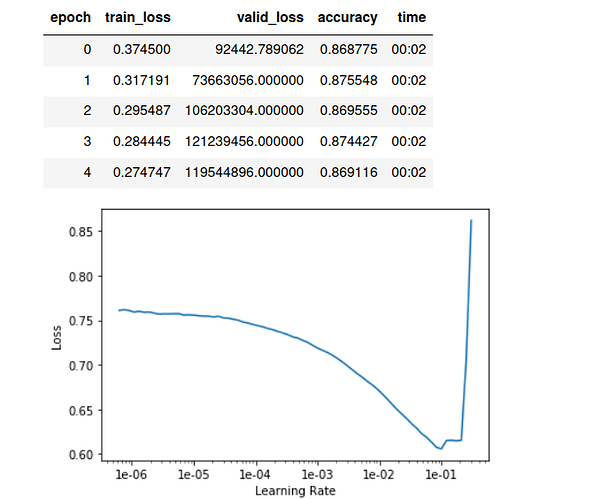
learn.lr_find()
learn.recorder.plot()
learn.fit_one_cycle(15, 1e-2, wd=0.01)
I also tried with these learning rates: 1e-3, 1e-2 and, 5e-3.
what is weird: loss is very high, while accuracy is reasonable.
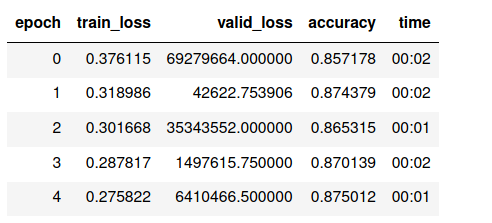
What I have investigated:
learn.loss_func returns:
FlattenedLoss of CrossEntropyLoss()
Is the model very naive and always predicting 0?
No; the validation set is 80% zeros and 20% ones. Also here is the CM:
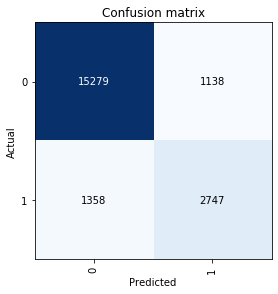
Is the architecture too big or too small?
I tried [100, 500], [10, 100, 10], [10, 10, 10], [10, 10], [10], [1000, 100]
All yielded very similar results, which is weird, isn’t it?
I tried with and without weight decay of 0.01, with and without emb_drop of 0.1 and 0.04, with and without ps=[0.001,0.01]
A random forests model had an accuracy of 87~88% and a boosted one reached 89% with close confusions matrices to what’s shown above.
What do you think is going on here?