Hi all,
I’ve developed a new optimizer, DeepMemory, that is testing out quite well so far.
Details:
DeepMemory is a new optimizer I came up with after blending DiffGrad + AdaMod. It takes the memory concept of AdaMod from merely throttling to actively blending and matches it to the epoch size.
The core concept is to provide the optimizer with long term memory of the previous step sizes across the entire epoch.
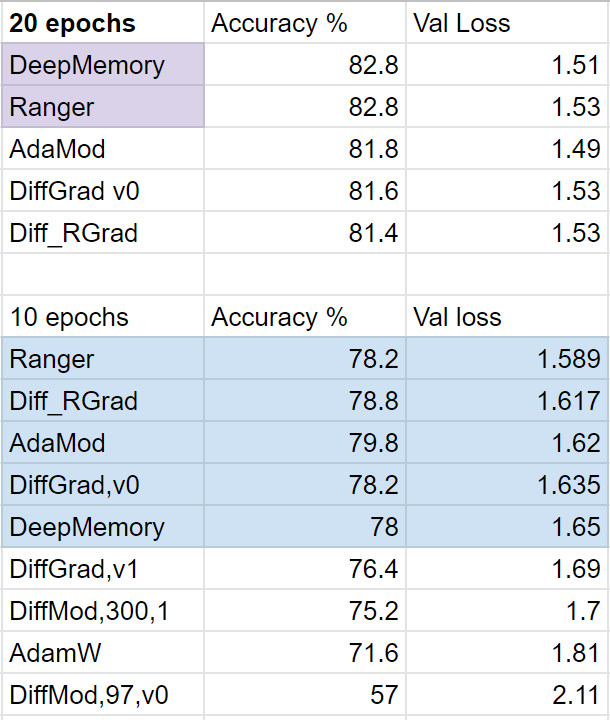
Results in initial testing put it on par with Ranger. Both Ranger and DeepMemory topped the recent testing I did with about 8 different optimizers (to be sure this is not conclusive! Just what I could do in an afternoon of testing and spending $20 on GPU time).
DeepMemory is designed to offset the weakness of many adaptive optimizers by creating a ‘long term’ memory of the gradients over the course of an epoch. This long term memory is averaged against the current adaptive step size generated from the current mini-batch in order to help guide the step size more optimally.
DeepMemory also keeps a short term gradient buffer that was developed in diffgrad, and locks down the step size when minimal gradient change is detected.
I’ll write up a more detailed article on Medium but for now if anyone is interested in testing, here’s the link along with a notebook already setup for running:
Usage:
DeepMemory works best when it knows the batches per epoch in order to match the memory size to your epochs:
memory_size = (len(data.x)//bs);memory_size #should be equal to or close to # of batches per epoch in order to build an average step size for the dataset
optar = partial(DeepMemory, len_memory = memory_size)
*please note you do need to copy over the deepmemory.py file to your working directory and then import.
from deepmemory import DeepMemory
Any feedback or questions are welcome. I’ll try and write up a more proper article soon.
Less