@fmobrj75 how are you declaring it on your Learner? (And the loss function)
Hi, @muellerzr
learn_lm.loss_func = LabelSmoothingCrossEntropy
Should it be different?
In fact I am trying to replicate the results of the paper https://arxiv.org/abs/1909.04761. I was able to replicate the results using fastai v1. But with fastai v2 the loss does not decrease with the same speed. I suspect it has something to do with the new format for the moms attributes and I want to use LabelSmoothingCrossEntropy when finetuning the language model on IMDB but receive this error.
Yes. It should be LabelSmoothingCrossEntropy()
Also would be very interested to see your notebook once you have finished it!  (I plan on doing Multi-FiT in the study group)
(I plan on doing Multi-FiT in the study group)
Thanks, @muellerzr.
I will set a github notebook set with it (fastai v1 and fastai2). I just have to solve the differences I am facing in fastai2 performance.
For the fastai v1, I followed @pierreguillou implementation based on the Multi-FIT paper.
When my experiments are set, I will notify you.
I tried using () and now I am receiving the following error:
ValueError: Expected target size (50, 15000), got torch.Size([50, 72])
Thanks that is much clearer now.
I created a LabelSmoothingCrossEntropyFlat inspired in the CrossEntropyLossFlat code and it is now working:
class LabelSmoothingCrossEntropyFlat(BaseLoss):
"Same as `nn.CrossEntropyLoss`, but flattens input and target."
y_int = True
def __init__(self, *args, axis=-1, **kwargs): super().__init__(LabelSmoothingCrossEntropy, *args, axis=axis, **kwargs)
def decodes(self, x): return x.argmax(dim=self.axis)
def activation(self, x): return F.softmax(x, dim=self.axis)@sgugger Managed to train a QRNN (1550 hidden and seq_len 70) language model and save model and encoder. When trying to train a QRNN classifier I got an error:
---------------------------------------------------------------------------
RuntimeError Traceback (most recent call last)
<ipython-input-58-d81c6bd29d71> in <module>
----> 1 learn.lr_find()
/media/hdd3tb/data/fastai2/fastai2/callback/schedule.py in lr_find(self, start_lr, end_lr, num_it, stop_div, show_plot)
194 n_epoch = num_it//len(self.dls.train) + 1
195 cb=LRFinder(start_lr=start_lr, end_lr=end_lr, num_it=num_it, stop_div=stop_div)
--> 196 with self.no_logging(): self.fit(n_epoch, cbs=cb)
197 if show_plot: self.recorder.plot_lr_find()
/media/hdd3tb/data/fastai2/fastai2/learner.py in fit(self, n_epoch, lr, wd, cbs, reset_opt)
287 try:
288 self.epoch=epoch; self('begin_epoch')
--> 289 self._do_epoch_train()
290 self._do_epoch_validate()
291 except CancelEpochException: self('after_cancel_epoch')
/media/hdd3tb/data/fastai2/fastai2/learner.py in _do_epoch_train(self)
262 try:
263 self.dl = self.dls.train; self('begin_train')
--> 264 self.all_batches()
265 except CancelTrainException: self('after_cancel_train')
266 finally: self('after_train')
/media/hdd3tb/data/fastai2/fastai2/learner.py in all_batches(self)
240 def all_batches(self):
241 self.n_iter = len(self.dl)
--> 242 for o in enumerate(self.dl): self.one_batch(*o)
243
244 def one_batch(self, i, b):
/media/hdd3tb/data/fastai2/fastai2/learner.py in one_batch(self, i, b)
246 try:
247 self._split(b); self('begin_batch')
--> 248 self.pred = self.model(*self.xb); self('after_pred')
249 if len(self.yb) == 0: return
250 self.loss = self.loss_func(self.pred, *self.yb); self('after_loss')
~/anaconda3/envs/fastai2/lib/python3.7/site-packages/torch/nn/modules/module.py in __call__(self, *input, **kwargs)
539 result = self._slow_forward(*input, **kwargs)
540 else:
--> 541 result = self.forward(*input, **kwargs)
542 for hook in self._forward_hooks.values():
543 hook_result = hook(self, input, result)
~/anaconda3/envs/fastai2/lib/python3.7/site-packages/torch/nn/modules/container.py in forward(self, input)
90 def forward(self, input):
91 for module in self._modules.values():
---> 92 input = module(input)
93 return input
94
~/anaconda3/envs/fastai2/lib/python3.7/site-packages/torch/nn/modules/module.py in __call__(self, *input, **kwargs)
539 result = self._slow_forward(*input, **kwargs)
540 else:
--> 541 result = self.forward(*input, **kwargs)
542 for hook in self._forward_hooks.values():
543 hook_result = hook(self, input, result)
/media/hdd3tb/data/fastai2/fastai2/text/models/core.py in forward(self, input)
92 #Note: this expects that sequence really begins on a round multiple of bptt
93 real_bs = (input[:,i] != self.pad_idx).long().sum()
---> 94 r,o = self.module(input[:real_bs,i: min(i+self.bptt, sl)])
95 if self.max_len is None or sl-i <= self.max_len:
96 raws.append(r)
~/anaconda3/envs/fastai2/lib/python3.7/site-packages/torch/nn/modules/module.py in __call__(self, *input, **kwargs)
539 result = self._slow_forward(*input, **kwargs)
540 else:
--> 541 result = self.forward(*input, **kwargs)
542 for hook in self._forward_hooks.values():
543 hook_result = hook(self, input, result)
/media/hdd3tb/data/fastai2/fastai2/text/models/awdlstm.py in forward(self, inp, from_embeds)
100 new_hidden,raw_outputs,outputs = [],[],[]
101 for l, (rnn,hid_dp) in enumerate(zip(self.rnns, self.hidden_dps)):
--> 102 raw_output, new_h = rnn(raw_output, self.hidden[l])
103 new_hidden.append(new_h)
104 raw_outputs.append(raw_output)
~/anaconda3/envs/fastai2/lib/python3.7/site-packages/torch/nn/modules/module.py in __call__(self, *input, **kwargs)
539 result = self._slow_forward(*input, **kwargs)
540 else:
--> 541 result = self.forward(*input, **kwargs)
542 for hook in self._forward_hooks.values():
543 hook_result = hook(self, input, result)
/media/hdd3tb/data/fastai2/fastai2/text/models/qrnn.py in forward(self, inp, hid)
152 if self.bidirectional: inp_bwd = inp.clone()
153 for i, layer in enumerate(self.layers):
--> 154 inp, h = layer(inp, None if hid is None else hid[2*i if self.bidirectional else i])
155 new_hid.append(h)
156 if self.bidirectional:
~/anaconda3/envs/fastai2/lib/python3.7/site-packages/torch/nn/modules/module.py in __call__(self, *input, **kwargs)
539 result = self._slow_forward(*input, **kwargs)
540 else:
--> 541 result = self.forward(*input, **kwargs)
542 for hook in self._forward_hooks.values():
543 hook_result = hook(self, input, result)
/media/hdd3tb/data/fastai2/fastai2/text/models/qrnn.py in forward(self, inp, hid)
100
101 def forward(self, inp, hid=None):
--> 102 y = self.linear(self._get_source(inp))
103 if self.output_gate: z_gate,f_gate,o_gate = y.chunk(3, dim=2)
104 else: z_gate,f_gate = y.chunk(2, dim=2)
/media/hdd3tb/data/fastai2/fastai2/text/models/qrnn.py in _get_source(self, inp)
123 if self.backward: inp_shift.insert(0,inp[:,1:] if self.batch_first else inp[1:])
124 else: inp_shift.append(inp[:,:-1] if self.batch_first else inp[:-1])
--> 125 inp_shift = torch.cat(inp_shift, dim)
126 return torch.cat([inp, inp_shift], 2)
127
RuntimeError: invalid argument 0: Sizes of tensors must match except in dimension 1. Got 2 and 1 in dimension 0 at /opt/conda/conda-bld/pytorch_1573049306803/work/aten/src/THC/generic/THCTensorMath.cu:71
The code that generated this error:
config = awd_qrnn_clas_config.copy()
config['n_hid'] = 1550
print(config)
{'emb_sz': 400,
'n_hid': 1550,
'n_layers': 4,
'pad_token': 1,
'bidir': False,
'output_p': 0.4,
'hidden_p': 0.3,
'input_p': 0.4,
'embed_p': 0.05,
'weight_p': 0.5}
learn = text_classifier_learner(dbunch,
AWD_QRNN,
pretrained=False,
config=config,
metrics=[accuracy],
path=path,
drop_mult=0.3,
loss_func=CrossEntropyLossFlat())
learn = learn.load_encoder('fastai2_15k_qrnn_multifit_fwd_enc')
learn.lr_find()
The same code with AWD_LSTM using an LSTM encoder works flawlessly.
I haven’t tested QRNNs thoroughly, so there may be bugs left in the code. I’ll look at this one when I have time (fair warning, we are on a book deadline so it might not be before a few weeks).
Ok. Thanks!
PR submitted to fastcore that should allow nested delegation.
Edit: I now seem to be getting an error in one of my fastai audio nbs relating to delegates. During import of fastai2.vision.augment I get this traceback, so it appears that my changes did break something and that I messed up the editable install step.
---------------------------------------------------------------------------
KeyError Traceback (most recent call last)
<ipython-input-12-9dd66b55d7bf> in <module>
----> 1 from fastai2.vision.augment import *
~/rob/fastai2/fastai2/vision/augment.py in <module>
11 from ..data.all import *
12 from .core import *
---> 13 from .data import *
14
15 # Cell
~/rob/fastai2/fastai2/vision/data.py in <module>
91
92 ImageDataBunch.from_csv = delegates(to=ImageDataBunch.from_df)(ImageDataBunch.from_csv)
---> 93 ImageDataBunch.from_path_re = delegates(to=ImageDataBunch.from_path_func)(ImageDataBunch.from_path_re)
94
95 # Cell
/opt/anaconda3/lib/python3.7/site-packages/fastcore/foundation.py in _f(f)
127 sig = inspect.signature(from_f)
128 sigd = dict(sig.parameters)
--> 129 k = sigd.pop('kwargs')
130 s2 = {k:v for k,v in inspect.signature(to_f).parameters.items()
131 if v.default != inspect.Parameter.empty and k not in sigd}
KeyError: 'kwargs'
The code that my change breaks is itself a nested delegation.
ImageDataBunch.from_path_re delegated to ImageDataBunch.from_path_func which delegates to DataBunch.from_dblock which delegates to TfmdDL.__init__.
Nested delegation without keep=True results in kwargs getting popped and not replaced which causes this error. I’m still not sure what the intention of these lines are, but possible fix would be
if keep: sigd['kwargs'] = k
else: from_f.__delwrap__ = to_f
from_f.__signature__ = sig.replace(parameters=sigd.values())
I’ll try it and update PR
Pushed update to the PR with solution proposed above. All appears to be working now but it could use a second set of eyes as I’m not sure I follow exactly what is happening at all points in delegates. Thank you and please let me know if there’s any additional ways I can help.
Quick question: is there any difference between:
learn(some_params).to_fp16()
and
learn(some_params, cbs=MixedPrecision())
Yes, there is another callback called ModelToHalf to add 
@sgugger I noticed many things can now be done with the DataLoader and the new args.
For example, we can now make partial epochs (for very large datasets) with very few changes (just get_idxs and shuffle_fn (for number of draws).
It will depend on the purpose of n arg. An idea would be to define an epoch as the following:
- if shuffle=False and
nis not set, we have full dataset (current implementation) - if shuffle=True and
nis not set, we have shuffled dataset (current implementation) - if shuffle=False and
nis set, we have a partial dataset (seems to be current implementation) - if shuffle=True and
nis set, we currently have a partial dataset that is shuffled, but we could do better since we loaded the full dataset. Why not returnnrandom draws from the full dataset?
If you think it’s a good idea I can also look at these changes which I think will be minimal.
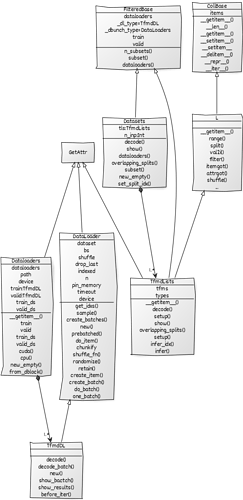
I would like to share with you a preliminary diagram of some fastai v2 classes that may be helpful to navigate between those classes. I created the diagram using VSCode and the yUML extension. You can also create it online. You can have a better preview and edit my file here. The diagram can be exported as png, jpg, pdf, or svg. Since it was just a proof of concept, I didn’t add all the information for those classes but if you spot any mistake, please let me know.
If you check the editable version you will notice it is a simple text file that I manually created using the following syntax:
[Customer]->[Order] // Association
[Customer]<>->[Order] // Aggregation
[Customer]++->[Order] // Composition
[Customer]1-0..1>[Order] // Cardinality
[Customer]1-0..orders 1>[Order] // Assoc Labels
[Customer]-.-[note: DAO] // Notes
[Customer]^[Member] // Inheritance
[Customer|name;address|save()] // Properties
Just a thought. I think it would be possible to add a custom function in nbdev to parse a jupyter notebook and automatically generate the corresponding .yuml file. One can start by parsing the Class definition and the doc section, and later add the class variables. For example:
class Datasets(FilteredBase):
...
_docs=dict(
decode="Compose `decode` of all `tuple_tfms` then all `tfms` on `i`",
show="Show item `o` in `ctx`",
dataloaders="Get a `DataLoaders`",
overlapping_splits="All splits that are in more than one split",
subset="New `Datasets` that only includes subset `i`",
new_empty="Create a new empty version of the `self`, keeping only the transforms",
set_split_idx="Contextmanager to use the same `Datasets` with another `split_idx`"
)
The result will be something like this:
[FilteredBase]^[Datasets|tls:TfmdLists;n_inp:Int|decode();show();dataloaders();overlapping_splits();subset();new_empty();set_split_idx()]
If you have a better idea on how to automatically generate UML with python, please let me know.
The last bit is unfortunate but changing it would change the meaning of the n attribute of DataLoader (which is the length of the underlying dataset) would change meaning, so I think it should be done with a custom get_idxs. We can add a function that does it easily for the user I guess.
Finished DataBlock.summary. A lot of folks complained it was rather hard to debug a data block, so you can now see everything that goes on when creating one.
First it tries to access one element of the dataset from a raw item, printing the results of every type transform you have (those are given by the blocks you use as well as your get_x and get_y functions).
Then it tries to create one sample, applying the item_transforms to the tuple you get from your dataset.
And finally it tries to collate a few samples in a batch (with a useful error message if that fails) before applying the batch transforms one at a time.
If dblock.summary(source) does not return any error, you have a verbose summary of all your input pipeline, and if it does, you know exactly at which points it failed, with hopefully all the information you need to fix the problem.
@sgugger I am really impressed with what you and jeremy are achieving. Using fastai 2 tweaks I was able to beat all my personal SOTAs with NLP Classification. My best results with fastai2 also beat all my best tries using huggingface transformers for all models I tried (bert mlm, roberta and xlm-roberta). Congratulations and thanks for your superb work!