I also faced the same problem, but don’t remember the exact reason for this. Look for it on keras github issues.
Try this:
for layer in fc_model.layers: layer.called_with = None conv_model.add(layer)
I also faced the same problem, but don’t remember the exact reason for this. Look for it on keras github issues.
Try this:
for layer in fc_model.layers: layer.called_with = None conv_model.add(layer)
Thank you @Manoj , it worked ! Although I don’t know why it raised this error , but it works, thanks again.
Hi, Jeremy,
In the lesson 3 notes, as to “filters”, there is a sentence “Then we would imagine that the brightest pixels in this new image are ones in a 3x3 area where the row above it s all 0 (black), and the center row is all 1 (white)”. How to understand it? I can’t catch what it means.
Thanks
Liu Peng
Hello everyone!
I’m having memory issues on lesson 3. The execution of the code runs fine until here:
conv_model.fit_generator(batches, samples_per_epoch=batches.nb_sample, nb_epoch=8,
validation_data=val_batches, nb_val_samples=val_batches.nb_sample)
After doing some stuff, I executed that line, getting a memory error after a couple of seconds. I tried re-executing all the code from scratch after a fresh reboot, and the program started fitting, but on the fourth epoch or so, I got the memory error again, which is the following:
MemoryError Traceback (most recent call last)
in ()
1 conv_model.fit_generator(batches, samples_per_epoch=batches.nb_sample, nb_epoch=8,
----> 2 validation_data=val_batches, nb_val_samples=val_batches.nb_sample)
/home/user/anaconda2/lib/python2.7/site-packages/keras/models.pyc in fit_generator(self, generator, samples_per_epoch, nb_epoch, verbose, callbacks, validation_data, nb_val_samples, class_weight, max_q_size, nb_worker, pickle_safe, initial_epoch, **kwargs)
933 nb_worker=nb_worker,
934 pickle_safe=pickle_safe,
–> 935 initial_epoch=initial_epoch)
936
937 def evaluate_generator(self, generator, val_samples,
/home/user/anaconda2/lib/python2.7/site-packages/keras/engine/training.pyc in fit_generator(self, generator, samples_per_epoch, nb_epoch, verbose, callbacks, validation_data, nb_val_samples, class_weight, max_q_size, nb_worker, pickle_safe, initial_epoch)
1555 outs = self.train_on_batch(x, y,
1556 sample_weight=sample_weight,
-> 1557 class_weight=class_weight)
1558
1559 if not isinstance(outs, list):
/home/user/anaconda2/lib/python2.7/site-packages/keras/engine/training.pyc in train_on_batch(self, x, y, sample_weight, class_weight)
1318 ins = x + y + sample_weights
1319 self._make_train_function()
-> 1320 outputs = self.train_function(ins)
1321 if len(outputs) == 1:
1322 return outputs[0]
/home/user/anaconda2/lib/python2.7/site-packages/keras/backend/theano_backend.pyc in call(self, inputs)
957 def call(self, inputs):
958 assert isinstance(inputs, (list, tuple))
–> 959 return self.function(*inputs)
960
961
/home/user/anaconda2/lib/python2.7/site-packages/theano/compile/function_module.pyc in call(self, *args, **kwargs)
896 node=self.fn.nodes[self.fn.position_of_error],
897 thunk=thunk,
–> 898 storage_map=getattr(self.fn, ‘storage_map’, None))
899 else:
900 # old-style linkers raise their own exceptions
/home/user/anaconda2/lib/python2.7/site-packages/theano/gof/link.pyc in raise_with_op(node, thunk, exc_info, storage_map)
323 # extra long error message in that case.
324 pass
–> 325 reraise(exc_type, exc_value, exc_trace)
326
327
/home/user/anaconda2/lib/python2.7/site-packages/theano/compile/function_module.pyc in call(self, *args, **kwargs)
882 try:
883 outputs =
–> 884 self.fn() if output_subset is None else
885 self.fn(output_subset=output_subset)
886 except Exception:
MemoryError: Error allocating 411041792 bytes of device memory (out of memory).
Apply node that caused the error: GpuAllocEmpty(Assert{msg=‘The convolution would produce an invalid shape (dim[0] < 0).’}.0, Assert{msg=‘The convolution would produce an invalid shape (dim[1] < 0).’}.0, Assert{msg=‘The convolution would produce an invalid shape (dim[2] <= 0).’}.0, Assert{msg=‘The convolution would produce an invalid shape (dim[3] <= 0).’}.0)
Toposort index: 211
Inputs types: [TensorType(int64, scalar), TensorType(int64, scalar), TensorType(int64, scalar), TensorType(int64, scalar)]
Inputs shapes: [(), (), (), ()]
Inputs strides: [(), (), (), ()]
Inputs values: [array(32), array(64), array(224), array(224)]
Outputs clients: [[GpuDnnConv{algo=‘small’, inplace=True}(GpuContiguous.0, GpuContiguous.0, GpuAllocEmpty.0, GpuDnnConvDesc{border_mode=‘valid’, subsample=(1, 1), conv_mode=‘conv’, precision=‘float32’}.0, Constant{1.0}, Constant{0.0})]]
HINT: Re-running with most Theano optimization disabled could give you a back-trace of when this node was created. This can be done with by setting the Theano flag ‘optimizer=fast_compile’. If that does not work, Theano optimizations can be disabled with ‘optimizer=None’.
HINT: Use the Theano flag ‘exception_verbosity=high’ for a debugprint and storage map footprint of this apply node.
I’m not sure where or why the error is produced… I’m running the course on my local machine, with 16 GB RAM and a GTX 970 running on Ubuntu.
Can someone help me? Thanks in advance.
Try reducing the batch size
Hi justinho,
That was bugging me as well. It turns out that updating the weights for bn_layers on line 80 updates the weights in the final model on line 79. I checked before and after just to be sure!
I could not find a compare weights function in Keras, so I use the code below to compare weights between 2 equally structured sequential models:
def compare_weights(model_1, model_2):
for i in range(len(model_1.layers)):
same = True
for j in range(len(model_1.layers[i].get_weights())):
if not (np.array_equal(model_1.layers[i].get_weights()[j], model_2.layers[i].get_weights()[j])):
same = False
print(same)thanks for sharing! 
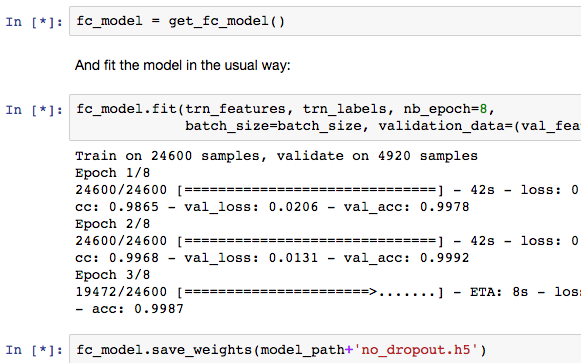
I had a question about the running notebooks. It has been my understanding that the * appears next to code while it is executing. However, during fitting sections of code it appears to have already executed (fc_model = get_fc_model) and still has the *. I believe the code is done executing because the fitting output from the code below (fc_model.fit) is showing. Usually after the fitting is complete the * corrects is replaced by a number.
Does anyone know why this is happening?
For example see this screenshot below.
My concern is more about debuggging. I have noticed the notebook hang when I execute several sections at the same time and Im not sure which section of the notebook needs attention.
I understand that overfitting first allows us to know the model is complex enough to handle the data.
My question is how to determine when we are overfitting. Just noting that val_acc is worse than training acc would seem to be not enough, in the case where Dropouts (applied to training but not validation) may artificially hamper training accuracy.
Should I train until val_acc begins to decrease instead, and not worry so much about comparing it directly to the training accuracy?
If validation is worse than training, then you’re overfitting - since dropout makes training worse, not better! However, you don’t want to have no overfitting - there’s some best amount. So you should focus on getting the best validation score you can, rather than the least overfitting.
In the notes about Zero-Padding, the first sentence reads:
… given that the filter necessarily operates on the premise that there are 8 surrounding pixels.
Why is the filter operating on the premise that there are 8 surrounding pixels? I thought it should be 9 surrounding pixels since the filter is a 3 x 3 matrix.
I’m confused about the role of vgg_ft(out_dim). My understanding is this updated model cuts off the last layer(s) and replaces it with a softmax layer with a specified number of outputs (2 in our case, dogs and cats). But Keras did this automatically for us using the Vgg16() model and our original 7 lines of code, right? Why do we have to hardcode this change now?
Hi @jeh0753,
In the original 7 lines of code, this was achieved by the following line:
vgg.finetune(batches)
finetune calls vgg_ft(batches.nb_class), which basically achieves the same thing.
@atk0 - Thanks for your reply. It looks to me as though vgg.finetune(batches) calls up vgg.ft(num), and vgg_ft(out_dim) is an unrelated alternative function that exists in utils.py. Do I have this right? And if so, why was vgg_ft(out_dim) built in the first place?
Hi again, Jake 
They’re not unrelated - the “global” vgg_ft(out_dim) function is simply a convenience function that creates a vgg16 model, performs finetuning on that model and then returns the model. Because part1 uses vgg16 frequently, Jeremy obviously decided that it would be nice to have a convenience function that does everything under the hood.
In the Lesson3,ipynb, I saw the In[6] is like this:
# Copy the weights from the pre-trained model.
# NB: Since we're removing dropout, we want to half the weights
def proc_wgts(layer): return [o/2 for o in layer.get_weights()]
This mean using the 1/2 of each value of weights, right? But dropout is related to reduce the number of weights, so I am confused why we divide each weight by 2 here. Can anyone help me understand it?
In addition, if the Dropout is not 0.5, but say, 0.8, do we still divide by 2?
How do I check the ‘summary’ of conv_layers and fc_layers after
conv_layers = layers[:last_conv_idx+1]
fc_layers = layers[last_conv_idx+1:]
They are lists, but I can only get something like this if I display the fc_layers:
[<keras.layers.pooling.MaxPooling2D at 0x7ff151153610>,
<keras.layers.core.Flatten at 0x7ff1510b64d0>,
<keras.layers.core.Dense at 0x7ff1511255d0>,
<keras.layers.core.Dropout at 0x7ff15111f850>,
<keras.layers.core.Dense at 0x7ff1510bf250>,
<keras.layers.core.Dropout at 0x7ff1510d9250>,
<keras.layers.core.Dense at 0x7ff1510ae290>]
How can I check the shape of each of these layers?? Thanks!
I have this question because when I do:
def get_fc_model():
model = Sequential([
MaxPooling2D(input_shape=conv_layers[-1].output_shape[1:]),
Flatten(),
Dense(4096, activation='relu'),
Dropout(0.),
Dense(4096, activation='relu'),
Dropout(0.),
Dense(3, activation='softmax')
])
for l1, l2 in zip(model.layers, fc_layers): l1.set_weights(proc_wgts(l2))
model.compile(optimizer=opt,
loss='categorical_crossentropy',
metrics=['accuracy'])
return model
fc_mdl = get_fc_model()
I got errors like:
ValueError Traceback (most recent call last)
in ()
----> 1 fc_mdl = get_fc_model()
in get_fc_model()
10 ])
11
—> 12 for l1, l2 in zip(model.layers, fc_layers): l1.set_weights(proc_wgts(l2))
13
14 model.compile(optimizer=opt,
/home/shi/anaconda2/lib/python2.7/site-packages/keras/engine/topology.pyc in set_weights(self, weights)
983 str(pv.shape) +
984 ’ not compatible with '
–> 985 'provided weight shape ’ + str(w.shape))
986 weight_value_tuples.append((p, w))
987 K.batch_set_value(weight_value_tuples)
ValueError: Layer weight shape (4096, 3) not compatible with provided weight shape (4096, 1000)
So I think the shape is wrong somewhere. Please help!
Hi, I have a question about training low level filters (that is, the first or second conv level). When we train the last level, we easily come up with loss function using log loss or simple sse. It’s based on the correct label and our predicted probability.
But for the first few levels, they are meant to capture abstract structures like edges and curves. Not only there’s no way to tell if the current activation map is correct (look at it manually??), but also it’s hard to get numerical loss (how wrong/right we are).
So, how do we train the set of weights (in this case, the values in the filter matrix) for the low level filters in practice?
Read some of the articles on back propagation. That’s how earlier layers get trained.
Hi,
First off apologies if this has been asked before. A search of the forums for State Farm didn’t turn up any related results. I am encountering issues with the State Farm data set. I am able to download the data set using the Kaggle cli tool after accepting the competitions terms. Checking the file sizes of the downloaded zip files confirms that I have correctly downloaded the data i.e. the imgs.zip file is 4.0GB on disk. I am unable to unzip the files. It appears that the zip files are corrupted in some way, or that they are not actually zip files.
unzip imgs.zip
results in the following output
Archive: imgs.zip
End-of-central-directory signature not found. Either this file is not
a zipfile, or it constitutes one disk of a multi-part archive. In the
latter case the central directory and zipfile comment will be found on
the last disk(s) of this archive.
unzip: cannot find zipfile directory in one of imgs.zip or
imgs.zip.zip, and cannot find imgs.zip.ZIP, period.
I also tried using the jar tool which was suggested on stackoverflow.
jar xf imgs.zip
Results in
Ick! 0
Myself and some others started the fast.ai course together recently. Others have encountered the same issue so it is definitely not isolated to my case. Has any one else encountered this issue and know of a solution or an alternate place to obtain the competition data set? Does this suggest that the competition has closed or something? I’m new to Kaggle.
Thanks,
Chris