Hi All,
I’m looking forward to the topics in part 2 of deep learning. While I think Jeremy’s notebooks are pretty self-explanatory this week, I’ve uploaded my notes into the forum as well. Hope its helpful!
- Tim
Welcome to Lesson 8
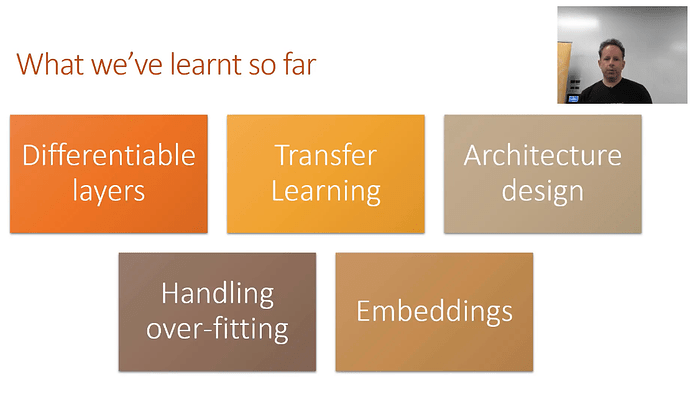
Where we are:
-
Differentiable Layers - coined by Yann Lecun, we don’t call it deep learning but differentiable programming. In part 1, we setup a diff’able function and a loss function that explains how good the parameters are. If you can define a loss function to score a task, and you have a flexible NN, you are done.
Yeah, Differentiable Programming is little more than a rebranding of the modern collection Deep Learning techniques, the same way Deep Learning was a rebranding of the modern incarnations of neural nets with more than two layers. The important point is that people are now building a new kind of software by assembling networks of parameterized functional blocks and by training them from examples using some form of gradient-based optimization….It’s really very much like a regular program, except it’s parameterized, automatically differentiated, and trainable/optimizable. Yann LeCun, Director of FAIR -
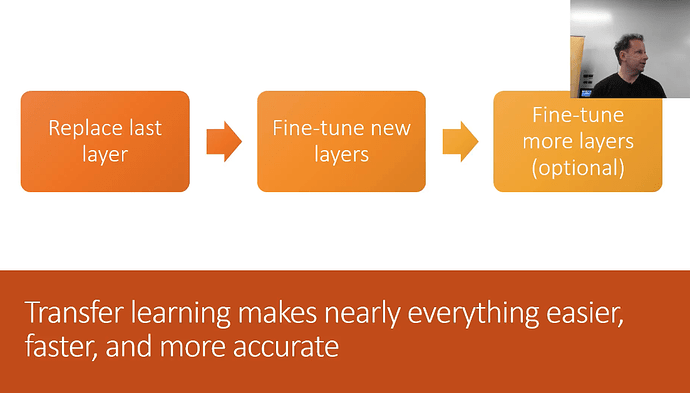
Transfer Learning - you should never need to start on random data. You should only start if no one has every solved a problem that is remotely close or related to what you are trying to solve. Fastai’s focus is completely focused on transfer learning and as a result the library is much different than any other library. In short, transfer learning takes a trained model, takes away the last few layers, retrains the last few layers, fine tunes the entire architecture, and as a result, trains a lot faster and requires less data!
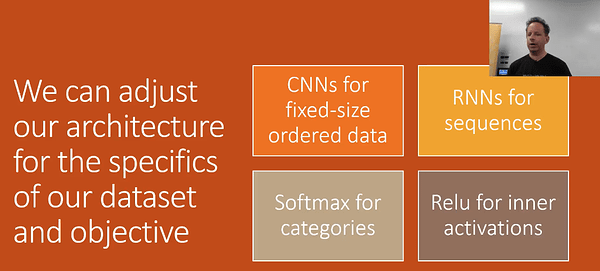
3.Architecture Design - There’s only a few models that generally cover a majority of problems. In part 1, we generally focused on activation functions and the output. We spend less time talking about architecture.
- how to Avoid overfitting - Create something that is over-parameterized. Train it for sure, overfit it, which guarantees that there is a predictive capability. Then we use the following techniques to reduce the overfitting to arrive at a generalized model.
- Get more data
- Data augmentation
- Generalized Architecture
- regularization
- Lastly, reduce architecture complexity
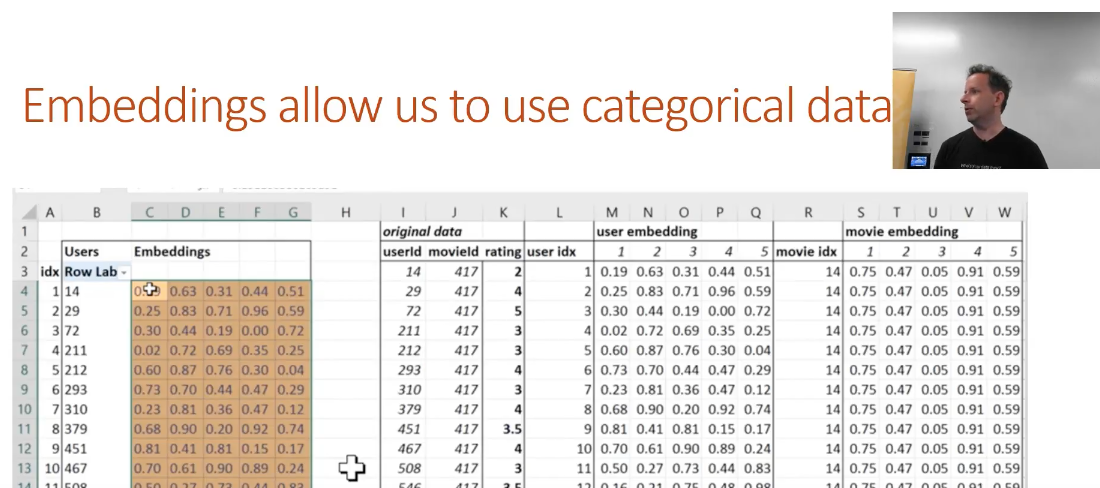
5.Embeddings - Just earlier last year, there was fewer discussions of using embeddings in tabular data (vs. the NLP related topics) Now these days there are more and more examples of using embeddings in traditional structured tabular data settings
Part 2’s Goals / Approach
Part 1 was an introduction to best practices. Overviewed mature techniques that were reasonably reliable for a wide variety of real world problems. These techniques were developed and tested over a longer period of time. Cutting Edge means that the best parameters may not always be evident. It may or may not be the absolute best solution, and the fastai implementation may still be buggy. The techniques covered will be promising in the research world, but may require tweaking. It’s exciting to work with the most current techniques and to also learn these recent techniques and understand what is going on vs. the recipe and pre-built libraries of pytorch and fastai.
Some caveats:
- Requires fastai customization
- Need to understand python well
- Other code resources will generally be research quality
- Code samples online nearly have always have problems
- Each lesson will be incomplete, ideas to explore
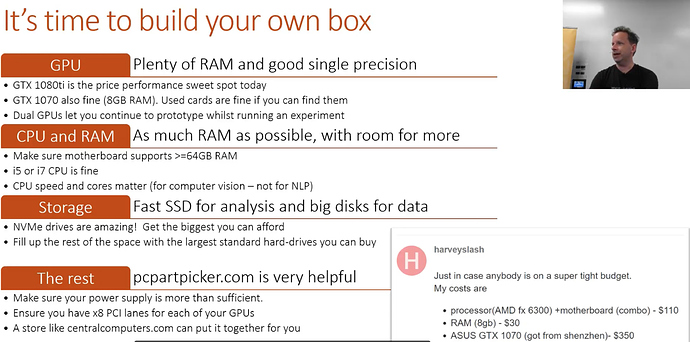
If you are considering building your own box checkout the forums.
It’s time to start reading papers
Each week we will be implementing a paper. In academic papers love using greek letters. Academics never refactor or substitute, but equations can get very long. Academic papers can be weird, but its the current way that research is commuted these days.
Since this is all cutting-edge, its a great opportunity!
- make a blog post, explain things in plain language
- maybe a simple implementation that other people can use
- use a published case study and translate the technique to a similar problem
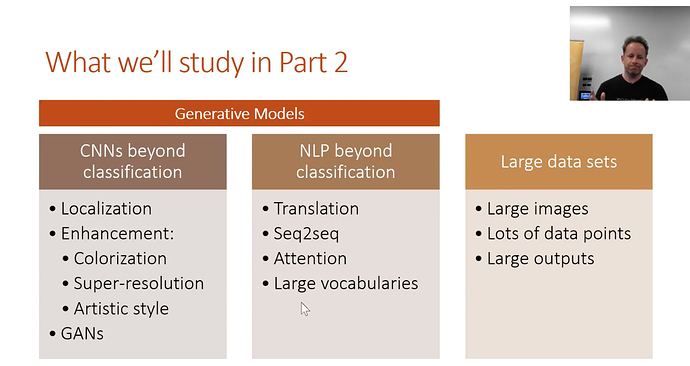
Part 2’s Topics
Generative models
NN’s generally output numbers. But now the outputs will be locations of objects, or a complete picture with a class in every pixel. Or an enhanced super-resolution of an image. Or text translated into french. Requires some different ways of thinking about things and different architectures.
Data: Text or Image data
Larger datasets - more objects and size of files
Object Detection
Introduction
1. We are classifying Multiple things
This part is not new, as we have done this in part1, the planet tutorial.
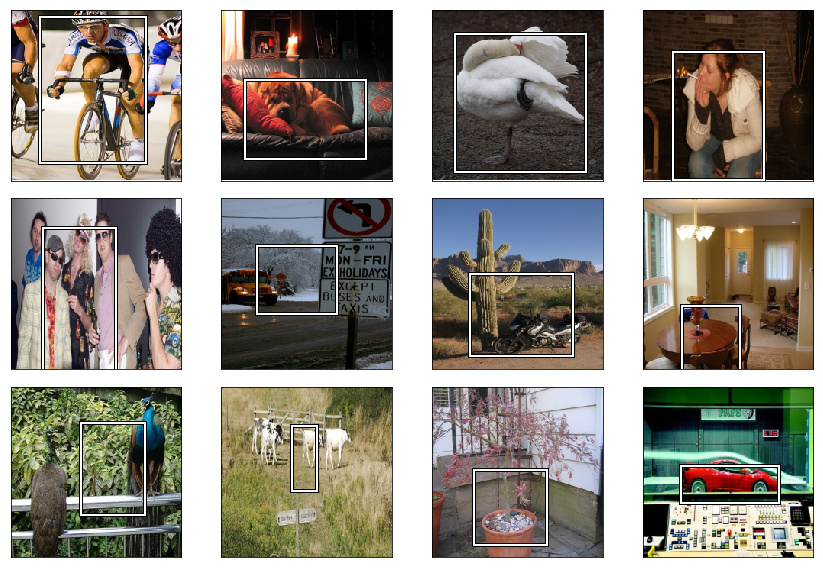
2. Finding Bounding boxes around the objects we are classifying
The box has the object entirely in it, but is no bigger than it needs to be. For these object detection datasets, we are looking for a pool of objects, but not necessarily EVERY object in the photo (mountain, tree cloud).
Stages:
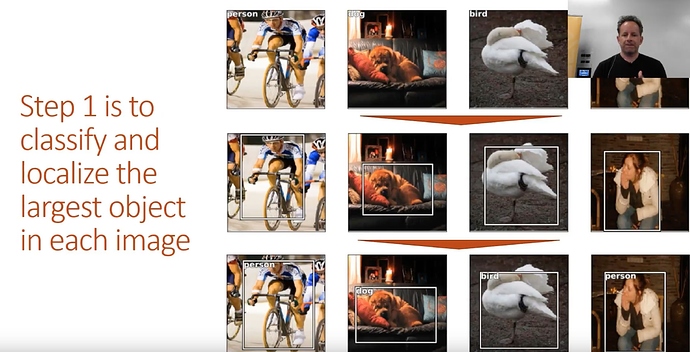
- 1 Find largest item
- 2 Find where it is
- 3 do both at the same time
Stage 1: Find the largest object in the image
Start with the Pascal Notebook
https://github.com/fastai/fastai/blob/master/courses/dl2/pascal.ipynb
%matplotlib inline
%reload_ext autoreload
%autoreload 2
Note, if you have a single GPU change the device to .set_device(0)
import sys
sys.path.append('../')
from fastai.conv_learner import *
from fastai.dataset import *
from pathlib import Path
import json
from PIL import ImageDraw, ImageFont
from matplotlib import patches, patheffects
# check to make sure you set the device
torch.cuda.set_device(0)
Pascal VOC
We will be looking at the Pascal VOC dataset. It’s quite slow, so you may prefer to download from this mirror. There are two different competition/research datasets, from 2007 and 2012. We’ll be using the 2007 version. You can use the larger 2012 for better results, or even combine them (but be careful to avoid data leakage between the validation sets if you do this).
Unlike previous lessons, we are using the python 3 standard library pathlib for our paths and file access. pathlib is the python3 library for interacting with files.
The pathlib module was first included in python 3.4 and has been enhanced in each of the subsequent releases. Pathlib is an object oriented interface to the filesystem and provides a more intuitive method to interact with the filesystem in a platform agnostic and pythonic manner.
Note that it returns an OS-specific class (on Linux, PosixPath) so your output may look a little different. Most libraries than take paths as input can take a pathlib object - although some (like cv2) can’t, in which case you can use str() to convert it to a string.
- Download the two zips
- make a
datadir - make a
pascaldir - move the
.jsonfiles out of the PASCAL_VOC into thepascal/dir
!wget http://pjreddie.com/media/files/VOCtrainval_06-Nov-2007.tar
--2018-03-21 01:00:51-- http://pjreddie.com/media/files/VOCtrainval_06-Nov-2007.tar
Resolving pjreddie.com (pjreddie.com)... 128.208.3.39
Connecting to pjreddie.com (pjreddie.com)|128.208.3.39|:80... connected.
HTTP request sent, awaiting response... 301 Moved Permanently
Location: https://pjreddie.com/media/files/VOCtrainval_06-Nov-2007.tar [following]
--2018-03-21 01:00:51-- https://pjreddie.com/media/files/VOCtrainval_06-Nov-2007.tar
Connecting to pjreddie.com (pjreddie.com)|128.208.3.39|:443... connected.
HTTP request sent, awaiting response... 200 OK
Length: 460032000 (439M) [application/octet-stream]
Saving to: ‘VOCtrainval_06-Nov-2007.tar’
VOCtrainval_06-Nov- 100%[===================>] 438.72M 12.7MB/s in 28s
2018-03-21 01:01:20 (15.5 MB/s) - ‘VOCtrainval_06-Nov-2007.tar’ saved [460032000/460032000]
!wget https://storage.googleapis.com/coco-dataset/external/PASCAL_VOC.zip
--2018-03-21 01:05:31-- https://storage.googleapis.com/coco-dataset/external/PASCAL_VOC.zip
Resolving storage.googleapis.com (storage.googleapis.com)... 172.217.6.48, 2607:f8b0:4005:805::2010
Connecting to storage.googleapis.com (storage.googleapis.com)|172.217.6.48|:443... connected.
HTTP request sent, awaiting response... 200 OK
Length: 1998182 (1.9M) [application/zip]
Saving to: ‘PASCAL_VOC.zip’
PASCAL_VOC.zip 100%[===================>] 1.91M --.-KB/s in 0.02s
2018-03-21 01:05:31 (77.5 MB/s) - ‘PASCAL_VOC.zip’ saved [1998182/1998182]
We will be using Python3’s path lib, creating a generator around the directory
PATH = Path('data/pascal')
list(PATH.iterdir())
[PosixPath('data/pascal/VOCdevkit'),
PosixPath('data/pascal/pascal_train2007.json'),
PosixPath('data/pascal/pascal_test2007.json'),
PosixPath('data/pascal/pascal_val2012.json'),
PosixPath('data/pascal/VOCtrainval_06-Nov-2007.tar'),
PosixPath('data/pascal/pascal_val2007.json'),
PosixPath('data/pascal/PASCAL_VOC.zip'),
PosixPath('data/pascal/pascal_train2012.json'),
PosixPath('data/pascal/PASCAL_VOC'),
PosixPath('data/pascal/models'),
PosixPath('data/pascal/src'),
PosixPath('data/pascal/tmp')]
Load the annotations
trn_j = json.load((PATH/'pascal_train2007.json').open())
trn_j.keys()
dict_keys(['images', 'type', 'annotations', 'categories'])
Image information
-
filename- the related image with filename -
height- how big the height of the image is -
width- how big the width of the image is -
id- the image id for joining to other datasets
IMAGES,ANNOTATIONS,CATEGORIES = ['images', 'annotations', 'categories']
trn_j[IMAGES][:5]
[{'file_name': '000012.jpg', 'height': 333, 'id': 12, 'width': 500},
{'file_name': '000017.jpg', 'height': 364, 'id': 17, 'width': 480},
{'file_name': '000023.jpg', 'height': 500, 'id': 23, 'width': 334},
{'file_name': '000026.jpg', 'height': 333, 'id': 26, 'width': 500},
{'file_name': '000032.jpg', 'height': 281, 'id': 32, 'width': 500}]
Bounding Boxes
-
bbox: column, row (top left) , height, width -
id: which image -
category_id: which label -
segmentation: ignore of this tutorail (polygon bounding
trn_j[ANNOTATIONS][:2]
[{'area': 34104,
'bbox': [155, 96, 196, 174],
'category_id': 7,
'id': 1,
'ignore': 0,
'image_id': 12,
'iscrowd': 0,
'segmentation': [[155, 96, 155, 270, 351, 270, 351, 96]]},
{'area': 13110,
'bbox': [184, 61, 95, 138],
'category_id': 15,
'id': 2,
'ignore': 0,
'image_id': 17,
'iscrowd': 0,
'segmentation': [[184, 61, 184, 199, 279, 199, 279, 61]]}]
Make a lookup from id to name
trn_j[CATEGORIES][:4]
[{'id': 1, 'name': 'aeroplane', 'supercategory': 'none'},
{'id': 2, 'name': 'bicycle', 'supercategory': 'none'},
{'id': 3, 'name': 'bird', 'supercategory': 'none'},
{'id': 4, 'name': 'boat', 'supercategory': 'none'}]
It’s helpful to use constants instead of strings, since we get tab-completion and don’t mistype.
FILE_NAME,ID,IMG_ID,CAT_ID,BBOX = 'file_name','id','image_id','category_id','bbox'
cats = dict((o[ID], o['name']) for o in trn_j[CATEGORIES])
trn_fns = dict((o[ID], o[FILE_NAME]) for o in trn_j[IMAGES])
trn_ids = [o[ID] for o in trn_j[IMAGES]]
Lets take a look at whats in the VOC 2007 dataset
list((PATH/'VOCdevkit'/'VOC2007').iterdir())
[PosixPath('data/pascal/VOCdevkit/VOC2007/ImageSets'),
PosixPath('data/pascal/VOCdevkit/VOC2007/SegmentationObject'),
PosixPath('data/pascal/VOCdevkit/VOC2007/SegmentationClass'),
PosixPath('data/pascal/VOCdevkit/VOC2007/Annotations'),
PosixPath('data/pascal/VOCdevkit/VOC2007/JPEGImages')]
Store the Image path
JPEGS = 'VOCdevkit/VOC2007/JPEGImages'
Make a compound path
IMG_PATH = PATH/JPEGS
list(IMG_PATH.iterdir())[:5]
[PosixPath('data/pascal/VOCdevkit/VOC2007/JPEGImages/006948.jpg'),
PosixPath('data/pascal/VOCdevkit/VOC2007/JPEGImages/005796.jpg'),
PosixPath('data/pascal/VOCdevkit/VOC2007/JPEGImages/007006.jpg'),
PosixPath('data/pascal/VOCdevkit/VOC2007/JPEGImages/004693.jpg'),
PosixPath('data/pascal/VOCdevkit/VOC2007/JPEGImages/002279.jpg')]
Each image has a unique ID.
im0_d = trn_j[IMAGES][0]
im0_d[FILE_NAME],im0_d[ID]
('000012.jpg', 12)
A defaultdict is useful any time you want to have a default dictionary entry for new keys. Here we create a dict from image IDs to a list of annotations (tuple of bounding box and class id).
We convert VOC’s height/width into top-left/bottom-right, and switch x/y coords to be consistent with numpy.
We are swapping dimensions, to be consistent, numpy, ROWS x COLUMNS
Output
{
IMG_ID : (array(top_left_row, top_left_col, lower_right_row, lower_right_col), CAT_ID)
}
#initialize the default dictionary
trn_anno = collections.defaultdict(lambda:[])
# for each annotation
for o in trn_j[ANNOTATIONS]:
# if not ignore
if not o['ignore']:
# get the original bounding box information
bb = o[BBOX]
bb = np.array([bb[1], bb[0], bb[3]+bb[1]-1, bb[2]+bb[0]-1])
#
trn_anno[o[IMG_ID]].append((bb,o[CAT_ID]))
len(trn_anno)
2501
example 1
im0_a = im_a[0]; im0_a
(array([ 96, 155, 269, 350]), 7)
cats[7]
'car'
example 2
trn_anno[17]
[(array([ 61, 184, 198, 278]), 15), (array([ 77, 89, 335, 402]), 13)]
cats[15],cats[13]
('person', 'horse')
Create a short function that will change a boundbox to height & width to translate data
def bb_hw(a): return np.array([a[1],a[0],a[3]-a[1],a[2]-a[0]])
We will use fast.ai’s open_image library to create an image to view
im = open_image(IMG_PATH/im0_d[FILE_NAME])
Create a short function to show the image in the notebook
Matplotlib’s plt.subplots is a really useful wrapper for creating plots, regardless of whether you have more than one subplot. Note that Matplotlib has an optional object-oriented API which I think is much easier to understand and use (although few examples online use it!)
def show_img(im, figsize=None, ax=None):
if not ax: fig,ax = plt.subplots(figsize=figsize)
ax.imshow(im)
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
return ax
Create an outlining function
A simple but rarely used trick to making text visible regardless of background is to use white text with black outline, or visa versa. Here’s how to do it in matplotlib.
def draw_outline(o, lw):
o.set_path_effects([patheffects.Stroke(
linewidth=lw, foreground='black'), patheffects.Normal()])
Note that * in argument lists is the splat operator. In this case it’s a little shortcut compared to writing out b[-2],b[-1].
Make a rectangle for the bounding box
def draw_rect(ax, b):
patch = ax.add_patch(patches.Rectangle(b[:2], *b[-2:], fill=False, edgecolor='white', lw=2))
draw_outline(patch, 4)
Make a quick function to write the text label (for the category)
def draw_text(ax, xy, txt, sz=14):
text = ax.text(*xy, txt,
verticalalignment='top', color='white', fontsize=sz, weight='bold')
draw_outline(text, 1)
Let’s test out showing an image!
ax = show_img(im)
b = bb_hw(im0_a[0])
draw_rect(ax, b)
draw_text(ax, b[:2], cats[im0_a[1]])

Let’s make a function to show multiple objects on a single image
def draw_im(im, ann):
ax = show_img(im, figsize=(16,8))
for b,c in ann:
b = bb_hw(b)
draw_rect(ax, b)
draw_text(ax, b[:2], cats[c], sz=16)
def draw_idx(i):
im_a = trn_anno[i]
im = open_image(IMG_PATH/trn_fns[i])
print(im.shape)
draw_im(im, im_a)
draw_idx(17)
(364, 480, 3)
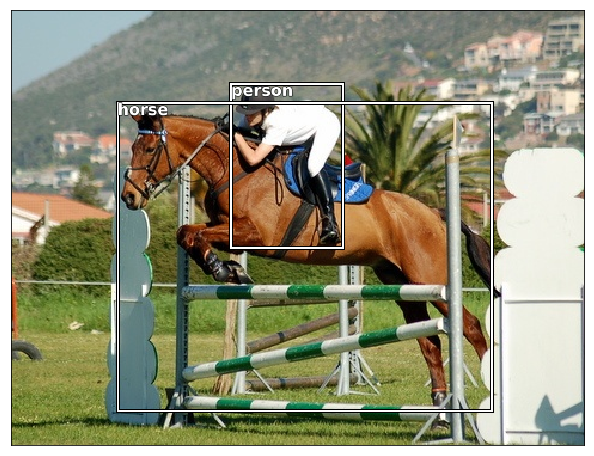

Next Complex step - Largest Item Classifier
A lambda function is simply a way to define an anonymous function inline. Here we use it to describe how to sort the annotation for each image - by bounding box size (descending).
This snippet sorts the objets
sorted(b, key=lambda x: np.product(x[0][-2:]-x[0][:2]), reverse=True)
We subtract the upper left from the bottom right and multiply (np.product) the values to get an area.
lambda x: np.product(x[0][-2:]-x[0][:2])
def get_lrg(b):
if not b: raise Exception()
b = sorted(b, key=lambda x: np.product(x[0][-2:]-x[0][:2]), reverse=True)
return b[0]
dictionary comprehension - storing the biggest objects:
{
IMG_ID : largest bounding box,
...
}
trn_lrg_anno = {a: get_lrg(b) for a,b in trn_anno.items()}
b,c = trn_lrg_anno[23]
b = bb_hw(b)
ax = show_img(open_image(IMG_PATH/trn_fns[23]), figsize=(5,10))
draw_rect(ax, b)
draw_text(ax, b[:2], cats[c], sz=16)

Let’s store the largest object per file in a CSV file
(PATH/'tmp').mkdir(exist_ok=True)
CSV = PATH/'tmp/lrg.csv'
Often it’s easiest to simply create a CSV of the data you want to model, rather than trying to create a custom dataset. Here we use Pandas to help us create a CSV of the image filename and class.
df = pd.DataFrame({'fn': [trn_fns[o] for o in trn_ids],
'cat': [cats[trn_lrg_anno[o][1]] for o in trn_ids]}, columns=['fn','cat'])
df.to_csv(CSV, index=False)
Make a Resnet model to learn
Considerations
- The training images are different sized
- Since we are doing bounding boxes, many of the objects are near the edge, so we want to be careful not to cut the objects off the picture
- So we will put
CropType.NOso we will “squish” instead
f_model = resnet34
sz=224
bs=64
From here it’s just like Dogs vs Cats!
tfms = tfms_from_model(f_model, sz, aug_tfms=transforms_side_on, crop_type=CropType.NO)
md = ImageClassifierData.from_csv(PATH, JPEGS, CSV, tfms=tfms)
A note on Data Loaders
A data loader is an iterator that will provide a mini-batch of data ( subset). But first we need to ensure that we start at the beginning of the dataset. Pythons’ iter() method will create an iterator object and start at the beginning of the dataset. And afterwards our iterator will have __next__ that can be used to pull a mini-batch
md.trn_dl
<fastai.dataloader.DataLoader at 0x7fba10572908>
Let’s look at a single example
x,y=next(iter(md.val_dl))
Some observations
- not a numpy array
- not on the CPU
-
[torch.cuda.FloatTensor of size 2x3x224x224 (GPU 0)]shape is wrong - not numbers between 0 and 1
x[:2]
( 0 , 0 ,.,.) =
3.9125e-01 4.3014e-01 4.8172e-01 ... 1.7518e-01 3.2367e-01 4.0783e-01
5.1636e-01 4.4973e-01 5.9202e-01 ... 1.7386e-01 2.3164e-01 3.6722e-01
5.4416e-01 5.7267e-01 7.0099e-01 ... 5.7682e-02 2.2320e-01 3.5455e-01
... ⋱ ...
1.4604e+00 1.5029e+00 1.5195e+00 ... 7.8030e-01 5.6716e-01 -6.3922e-01
9.3739e-01 1.0210e+00 1.1599e+00 ... 1.1281e+00 1.0895e+00 4.5857e-01
5.8584e-01 4.5245e-01 2.9605e-01 ... 1.0003e+00 9.2495e-01 8.2729e-01
( 0 , 1 ,.,.) =
2.4041e-01 3.1444e-01 4.1422e-01 ... 3.3162e-01 4.7052e-01 5.4764e-01
3.9737e-01 4.2156e-01 5.7304e-01 ... 3.3887e-01 3.8799e-01 5.2038e-01
5.2462e-01 5.8245e-01 6.7485e-01 ... 2.5190e-01 4.0003e-01 5.1502e-01
... ⋱ ...
1.4721e+00 1.5019e+00 1.5077e+00 ... 6.0917e-01 4.4337e-01 -7.3978e-01
8.4169e-01 9.4566e-01 1.0678e+00 ... 9.7373e-01 1.0164e+00 3.9674e-01
4.7731e-01 3.6442e-01 1.8230e-01 ... 8.5592e-01 8.5008e-01 7.5529e-01
( 0 , 2 ,.,.) =
6.3094e-01 7.6758e-01 9.1924e-01 ... 4.6997e-01 5.8218e-01 6.3571e-01
8.6685e-01 8.9343e-01 1.0892e+00 ... 4.5970e-01 4.9210e-01 6.0825e-01
9.6684e-01 1.0009e+00 1.0785e+00 ... 3.5109e-01 5.0631e-01 6.3579e-01
... ⋱ ...
1.5606e+00 1.5927e+00 1.6055e+00 ... 5.4531e-01 3.8118e-01 -7.9327e-01
9.2625e-01 1.0206e+00 1.1445e+00 ... 9.1826e-01 9.3747e-01 3.1286e-01
5.6966e-01 4.4559e-01 2.6428e-01 ... 8.0080e-01 7.7093e-01 6.7405e-01
⋮
( 1 , 0 ,.,.) =
2.1343e+00 2.1323e+00 2.1205e+00 ... -2.0870e+00 -2.0843e+00 -2.0837e+00
2.1703e+00 2.1498e+00 2.1186e+00 ... -2.0939e+00 -2.0863e+00 -2.0861e+00
2.1454e+00 2.1375e+00 2.0940e+00 ... -2.0922e+00 -2.0923e+00 -2.0933e+00
... ⋱ ...
-1.8905e+00 -1.8818e+00 -1.8374e+00 ... -1.8581e+00 -1.8778e+00 -1.8109e+00
-1.8530e+00 -1.8759e+00 -1.8143e+00 ... -1.8267e+00 -1.8599e+00 -1.8636e+00
-1.8764e+00 -1.8944e+00 -1.9561e+00 ... -1.8202e+00 -1.7856e+00 -1.8230e+00
( 1 , 1 ,.,.) =
2.2305e+00 2.1392e+00 1.7919e+00 ... -2.0042e+00 -2.0013e+00 -2.0007e+00
2.2569e+00 2.1649e+00 1.8062e+00 ... -2.0112e+00 -2.0034e+00 -2.0032e+00
2.2407e+00 2.1826e+00 1.8103e+00 ... -2.0095e+00 -2.0096e+00 -2.0105e+00
... ⋱ ...
-1.8032e+00 -1.7944e+00 -1.7489e+00 ... -1.7557e+00 -1.7819e+00 -1.7218e+00
-1.7649e+00 -1.7883e+00 -1.7254e+00 ... -1.7107e+00 -1.7613e+00 -1.7757e+00
-1.7888e+00 -1.8072e+00 -1.8703e+00 ... -1.7174e+00 -1.6914e+00 -1.7343e+00
( 1 , 2 ,.,.) =
1.8575e+00 1.3323e+00 6.5693e-01 ... -1.7730e+00 -1.7702e+00 -1.7696e+00
1.9155e+00 1.3577e+00 6.7138e-01 ... -1.7800e+00 -1.7723e+00 -1.7721e+00
1.8913e+00 1.3744e+00 6.8420e-01 ... -1.7783e+00 -1.7784e+00 -1.7794e+00
... ⋱ ...
-1.5730e+00 -1.5642e+00 -1.5189e+00 ... -1.5329e+00 -1.5559e+00 -1.4919e+00
-1.5348e+00 -1.5581e+00 -1.4955e+00 ... -1.4945e+00 -1.5366e+00 -1.5456e+00
-1.5586e+00 -1.5769e+00 -1.6397e+00 ... -1.4945e+00 -1.4639e+00 -1.5043e+00
[torch.cuda.FloatTensor of size 2x3x224x224 (GPU 0)]
imagenet_stats = A([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
inception_stats = A([0.5, 0.5, 0.5], [0.5, 0.5, 0.5])
inception_models = (inception_4, inceptionresnet_2)
def tfms_from_stats(stats, sz, aug_tfms=None, max_zoom=None, pad=0, crop_type=CropType.RANDOM,
tfm_y=None, sz_y=None, pad_mode=cv2.BORDER_REFLECT):
if aug_tfms is None: aug_tfms=[]
----------------------------------------------- NOTE THE MANIPULATION ----------------------------------
tfm_norm = Normalize(*stats, tfm_y=tfm_y)
tfm_denorm = Denormalize(*stats)
----------------------------------------------- NOTE THE MANIPULATION ----------------------------------
val_crop = CropType.CENTER if crop_type==CropType.RANDOM else crop_type
val_tfm = image_gen(tfm_norm, tfm_denorm, sz, pad=pad, crop_type=val_crop, tfm_y=tfm_y, sz_y=sz_y)
trn_tfm=image_gen(tfm_norm, tfm_denorm, sz, tfms=aug_tfms, max_zoom=max_zoom,
pad=pad, crop_type=crop_type, tfm_y=tfm_y, sz_y=sz_y, pad_mode=pad_mode)
return trn_tfm, val_tfm
show_img(md.val_ds.denorm(to_np(x))[0]);

Using Resnet 34
Get a pretrained learner
learn = ConvLearner.pretrained(f_model, md, metrics=[accuracy])
learn.opt_fn = optim.Adam
Find a learning rate
lrf=learn.lr_find(1e-5,100)
Failed to display Jupyter Widget of type HBox.
If you're reading this message in the Jupyter Notebook or JupyterLab Notebook, it may mean that the widgets JavaScript is still loading. If this message persists, it likely means that the widgets JavaScript library is either not installed or not enabled. See the Jupyter Widgets Documentation for setup instructions.
If you're reading this message in another frontend (for example, a static rendering on GitHub or NBViewer), it may mean that your frontend doesn't currently support widgets.
78%|███████▊ | 25/32 [00:08<00:02, 2.84it/s, loss=14.9]
learn.sched.plot()
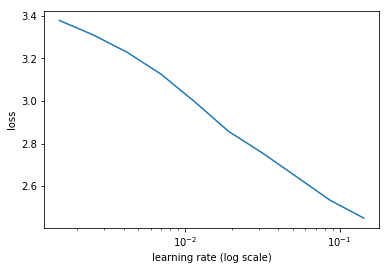
to change the truncation of the plot use the command below
learn.sched.plot(n_skip=5, n_skip_end=1)
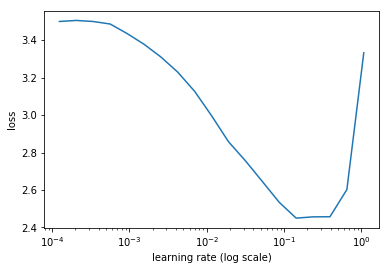
Set the learning rate
lr = 2e-2
learn.fit(lr, 1, cycle_len=1)
Failed to display Jupyter Widget of type HBox.
If you're reading this message in the Jupyter Notebook or JupyterLab Notebook, it may mean that the widgets JavaScript is still loading. If this message persists, it likely means that the widgets JavaScript library is either not installed or not enabled. See the Jupyter Widgets Documentation for setup instructions.
If you're reading this message in another frontend (for example, a static rendering on GitHub or NBViewer), it may mean that your frontend doesn't currently support widgets.
epoch trn_loss val_loss accuracy
0 1.280753 0.604127 0.806941
[0.60412693, 0.8069411069154739]
lrs = np.array([lr/1000,lr/100,lr])
We freeze all layers except the last two layers, find new learning rate and retrain
learn.freeze_to(-2)
lrf=learn.lr_find(lrs/1000)
learn.sched.plot(1)
Failed to display Jupyter Widget of type HBox.
If you're reading this message in the Jupyter Notebook or JupyterLab Notebook, it may mean that the widgets JavaScript is still loading. If this message persists, it likely means that the widgets JavaScript library is either not installed or not enabled. See the Jupyter Widgets Documentation for setup instructions.
If you're reading this message in another frontend (for example, a static rendering on GitHub or NBViewer), it may mean that your frontend doesn't currently support widgets.
84%|████████▍ | 27/32 [00:08<00:01, 3.26it/s, loss=3.45]
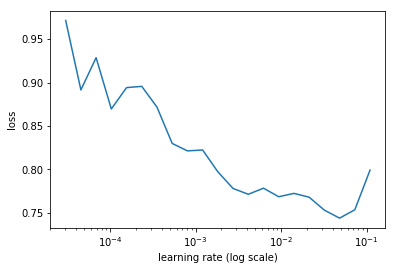
learn.fit(lrs/5, 1, cycle_len=1)
Failed to display Jupyter Widget of type HBox.
If you're reading this message in the Jupyter Notebook or JupyterLab Notebook, it may mean that the widgets JavaScript is still loading. If this message persists, it likely means that the widgets JavaScript library is either not installed or not enabled. See the Jupyter Widgets Documentation for setup instructions.
If you're reading this message in another frontend (for example, a static rendering on GitHub or NBViewer), it may mean that your frontend doesn't currently support widgets.
epoch trn_loss val_loss accuracy
0 0.777971 0.556964 0.832782
[0.5569643, 0.8327824547886848]
Accuracy is still at 83%
learn.unfreeze()
Accuracy isn’t improving much - since many images have multiple different objects, it’s going to be impossible to be that accurate.
learn.fit(lrs/5, 1, cycle_len=2)
Failed to display Jupyter Widget of type HBox.
If you're reading this message in the Jupyter Notebook or JupyterLab Notebook, it may mean that the widgets JavaScript is still loading. If this message persists, it likely means that the widgets JavaScript library is either not installed or not enabled. See the Jupyter Widgets Documentation for setup instructions.
If you're reading this message in another frontend (for example, a static rendering on GitHub or NBViewer), it may mean that your frontend doesn't currently support widgets.
epoch trn_loss val_loss accuracy
0 0.676254 0.546998 0.834285
1 0.460609 0.533741 0.833233
[0.53374064, 0.8332331702113152]
learn.save('clas_one')
learn.load('clas_one')
x,y = next(iter(md.val_dl))
probs = F.softmax(predict_batch(learn.model, x), -1)
x,preds = to_np(x),to_np(probs)
preds = np.argmax(preds, -1)
Let’s look at the 20 classes
fig, axes = plt.subplots(3, 4, figsize=(12, 8))
for i,ax in enumerate(axes.flat):
ima=md.val_ds.denorm(x)[i]
b = md.classes[preds[i]]
ax = show_img(ima, ax=ax)
draw_text(ax, (0,0), b)
plt.tight_layout()
Clipping input data to the valid range for imshow with RGB data ([0..1] for floats or [0..255] for integers).
Note on debuggers
You can use the python debugger pdb to step through code.
-
pdb.set_trace()to set a breakpoint -
%debugmagic to trace an error
Commands you need to know:
- s / n / c
- u / d
- p
- l
> <ipython-input-99-6d7dd6a3d3cc>(4)<module>()
-> ima=md.val_ds.denorm(x)[i]
(Pdb) h
Documented commands (type help <topic>):
========================================
EOF c d h list q rv undisplay
a cl debug help ll quit s unt
alias clear disable ignore longlist r source until
args commands display interact n restart step up
b condition down j next return tbreak w
break cont enable jump p retval u whatis
bt continue exit l pp run unalias where
Miscellaneous help topics:
==========================
exec pdb
Can view variables throughout the debugging process
(Pdb) n # will go to next step
(Pdb) l # will show the currentlocation
1 fig, axes = plt.subplots(3, 4, figsize=(12, 8))
2 for i,ax in enumerate(axes.flat):
3 pdb.set_trace()
4 -> ima=md.val_ds.denorm(x)[i]
5 b = md.classes[preds[i]]
6 ax = show_img(ima, ax=ax)
7 draw_text(ax, (0,0), b)
8 plt.tight_layout()
[EOF]
(Pdb) s # will go into a function
(Pdb) c # continue to next break point
Next Stage : Create a bounding box around an object
We know we can make a regression nn instead of a classification. This is accomplished by changing the last layer of the NN. Instead of Softmax, and use MSE, it is now a regression problem. We can have multiple outputs.
So what we will do is a multiple regression to predict the following values:
- top left x
- top left y
- lower right x
- lower right y
But what about the loss function?
BB_CSV = PATH/'tmp/bb.csv'
Transform the bounding box data
bb = np.array([trn_lrg_anno[o][0] for o in trn_ids])
bbs = [' '.join(str(p) for p in o) for o in bb]
df = pd.DataFrame({'fn': [trn_fns[o] for o in trn_ids], 'bbox': bbs}, columns=['fn','bbox'])
df.to_csv(BB_CSV, index=False)
BB_CSV.open().readlines()[:5]
['fn,bbox\n',
'000012.jpg,96 155 269 350\n',
'000017.jpg,77 89 335 402\n',
'000023.jpg,1 2 461 242\n',
'000026.jpg,124 89 211 336\n']
Set our model and parameters
f_model=resnet34
sz=224
bs=64
Tell fast.ai to make a continous network model
Set continuous=True to tell fastai this is a regression problem, which means it won’t one-hot encode the labels, and will use MSE as the default crit.
Note that we have to tell the transforms constructor that our labels are coordinates, so that it can handle the transforms correctly.
Also, we use CropType.NO because we want to ‘squish’ the rectangular images into squares, rather than center cropping, so that we don’t accidentally crop out some of the objects. (This is less of an issue in something like imagenet, where there is a single object to classify, and it’s generally large and centrally located).
tfms = tfms_from_model(f_model, sz, crop_type=CropType.NO, tfm_y=TfmType.COORD)
md = ImageClassifierData.from_csv(PATH, JPEGS, BB_CSV, tfms=tfms, continuous=True)
x,y=next(iter(md.val_dl))
ima=md.val_ds.denorm(to_np(x))[0]
b = bb_hw(to_np(y[0])); b
array([ 49., 0., 131., 205.], dtype=float32)
Let’s look at an example image with box
ax = show_img(ima)
draw_rect(ax, b)
draw_text(ax, b[:2], 'label')

How add additional layers on the end of the Resnet (custom head)
fastai let’s you use a custom_head to add your own module on top of a convnet, instead of the adaptive pooling and fully connected net which is added by default. In this case, we don’t want to do any pooling, since we need to know the activations of each grid cell.
The final layer has 4 activations, one per bounding box coordinate. Our target is continuous, not categorical, so the MSE loss function used does not do any sigmoid or softmax to the module outputs.
head_reg4 = nn.Sequential(Flatten(), nn.Linear(25088,4))
learn = ConvLearner.pretrained(f_model, md, custom_head=head_reg4)
learn.opt_fn = optim.Adam
learn.crit = nn.L1Loss()
Check the Model to see that the additional layer has been addedm
# learn.summary()
OrderedDict([('Conv2d-1',
OrderedDict([('input_shape', [-1, 3, 224, 224]),
('output_shape', [-1, 64, 112, 112]),
('trainable', False),
('nb_params', 9408)])),
....
....
('ReLU-121',
OrderedDict([('input_shape', [-1, 512, 7, 7]),
('output_shape', [-1, 512, 7, 7]),
('nb_params', 0)])),
('BasicBlock-122',
OrderedDict([('input_shape', [-1, 512, 7, 7]),
('output_shape', [-1, 512, 7, 7]),
('nb_params', 0)])),
('Flatten-123',
OrderedDict([('input_shape', [-1, 512, 7, 7]),
('output_shape', [-1, 25088]),
('nb_params', 0)])),
------------------- New layer ------------------------
('Linear-124',
OrderedDict([('input_shape', [-1, 25088]),
('output_shape', [-1, 4]),
('trainable', True),
('nb_params', 100356)]))])
------------------- New layer ------------------------
Try and fit the model
learn.lr_find(1e-5,100)
learn.sched.plot(5)
Failed to display Jupyter Widget of type HBox.
If you're reading this message in the Jupyter Notebook or JupyterLab Notebook, it may mean that the widgets JavaScript is still loading. If this message persists, it likely means that the widgets JavaScript library is either not installed or not enabled. See the Jupyter Widgets Documentation for setup instructions.
If you're reading this message in another frontend (for example, a static rendering on GitHub or NBViewer), it may mean that your frontend doesn't currently support widgets.
78%|███████▊ | 25/32 [00:04<00:01, 5.36it/s, loss=475]
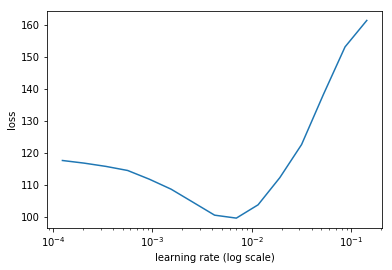
Set the learning rate
lr = 2e-3
Train
learn.fit(lr, 2, cycle_len=1, cycle_mult=2)
Failed to display Jupyter Widget of type HBox.
If you're reading this message in the Jupyter Notebook or JupyterLab Notebook, it may mean that the widgets JavaScript is still loading. If this message persists, it likely means that the widgets JavaScript library is either not installed or not enabled. See the Jupyter Widgets Documentation for setup instructions.
If you're reading this message in another frontend (for example, a static rendering on GitHub or NBViewer), it may mean that your frontend doesn't currently support widgets.
epoch trn_loss val_loss
0 50.135777 34.477402
1 37.689602 29.124092
2 31.387475 27.658106
[27.658106]
lrs = np.array([lr/100,lr/10,lr])
learn.freeze_to(-2)
lrf=learn.lr_find(lrs/1000)
learn.sched.plot(1)
Failed to display Jupyter Widget of type HBox.
If you're reading this message in the Jupyter Notebook or JupyterLab Notebook, it may mean that the widgets JavaScript is still loading. If this message persists, it likely means that the widgets JavaScript library is either not installed or not enabled. See the Jupyter Widgets Documentation for setup instructions.
If you're reading this message in another frontend (for example, a static rendering on GitHub or NBViewer), it may mean that your frontend doesn't currently support widgets.
epoch trn_loss val_loss
0 80.37384 175370041032704.0
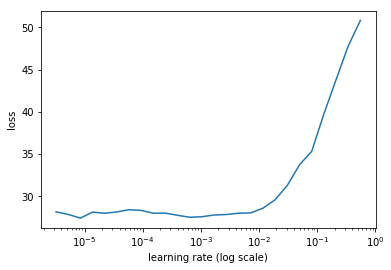
learn.fit(lrs, 2, cycle_len=1, cycle_mult=2)
Failed to display Jupyter Widget of type HBox.
If you're reading this message in the Jupyter Notebook or JupyterLab Notebook, it may mean that the widgets JavaScript is still loading. If this message persists, it likely means that the widgets JavaScript library is either not installed or not enabled. See the Jupyter Widgets Documentation for setup instructions.
If you're reading this message in another frontend (for example, a static rendering on GitHub or NBViewer), it may mean that your frontend doesn't currently support widgets.
epoch trn_loss val_loss
0 25.814654 23.127014
1 21.655237 21.125538
2 17.600573 20.209145
[20.209145]
learn.freeze_to(-3)
learn.fit(lrs, 1, cycle_len=2)
Failed to display Jupyter Widget of type HBox.
If you're reading this message in the Jupyter Notebook or JupyterLab Notebook, it may mean that the widgets JavaScript is still loading. If this message persists, it likely means that the widgets JavaScript library is either not installed or not enabled. See the Jupyter Widgets Documentation for setup instructions.
If you're reading this message in another frontend (for example, a static rendering on GitHub or NBViewer), it may mean that your frontend doesn't currently support widgets.
epoch trn_loss val_loss
0 16.644847 21.78323
1 14.667386 20.380457
[20.380457]
Save our Model
learn.save('reg4')
learn.load('reg4')
Let’s see how our model did!
Anytime there’s a single subject, our model does decent. When there’s multiple objects our model doesn’t perform as well. Next week we will improve our model
x,y = next(iter(md.val_dl))
learn.model.eval()
preds = to_np(learn.model(VV(x)))
fig, axes = plt.subplots(3, 4, figsize=(12, 8))
for i,ax in enumerate(axes.flat):
ima=md.val_ds.denorm(to_np(x))[i]
b = bb_hw(preds[i])
ax = show_img(ima, ax=ax)
draw_rect(ax, b)
plt.tight_layout()
Clipping input data to the valid range for imshow with RGB data ([0..1] for floats or [0..255] for integers).