My data is organised as:
train:- containing folder for each class
valid :- containing folder for each class
test:- contains all test images
I have the same issue.
Without seeing the entire error message, my first guess would be to delete the tmp folder and then try again.
I have the same issue with the lesson1 notebook. It works fine using the dogscats dataset, but when I use another dataset with dog breeds, it fails with the same error as above. Deleting the tmp directory doesn’t help. This is the entire stack trace:
---------------------------------------------------------------------------
AttributeError Traceback (most recent call last)
<ipython-input-17-42d5f498bf97> in <module>()
1 arch=resnet34
2 data = ImageClassifierData.from_paths(PATH, tfms=tfms_from_model(arch, sz))
----> 3 learn = ConvLearner.pretrained(arch, data, precompute=True)
4 learn.fit(0.01, 3)
~\fastai\fastai\conv_learner.py in pretrained(cls, f, data, ps, xtra_fc, xtra_cut, **kwargs)
96 def pretrained(cls, f, data, ps=None, xtra_fc=None, xtra_cut=0, **kwargs):
97 models = ConvnetBuilder(f, data.c, data.is_multi, data.is_reg, ps=ps, xtra_fc=xtra_fc, xtra_cut=xtra_cut)
---> 98 return cls(data, models, **kwargs)
99
100 @property
~\fastai\fastai\conv_learner.py in __init__(self, data, models, precompute, **kwargs)
89 elif self.metrics is None:
90 self.metrics = [accuracy_multi] if self.data.is_multi else [accuracy]
---> 91 if precompute: self.save_fc1()
92 self.freeze()
93 self.precompute = precompute
~\fastai\fastai\conv_learner.py in save_fc1(self)
135 m=self.models.top_model
136 if len(self.activations[0])!=len(self.data.trn_ds):
--> 137 predict_to_bcolz(m, self.data.fix_dl, act)
138 if len(self.activations[1])!=len(self.data.val_ds):
139 predict_to_bcolz(m, self.data.val_dl, val_act)
~\fastai\fastai\model.py in predict_to_bcolz(m, gen, arr, workers)
11 lock=threading.Lock()
12 m.eval()
---> 13 for x,*_ in tqdm(gen):
14 y = to_np(m(VV(x)).data)
15 with lock:
c:\users\hille\anaconda3\envs\dlwin36\lib\site-packages\tqdm\_tqdm.py in __iter__(self)
951 """, fp_write=getattr(self.fp, 'write', sys.stderr.write))
952
--> 953 for obj in iterable:
954 yield obj
955 # Update and possibly print the progressbar.
~\fastai\fastai\dataset.py in __next__(self)
242 if self.i>=len(self.dl): raise StopIteration
243 self.i+=1
--> 244 return next(self.it)
245
246 @property
~\fastai\fastai\dataloader.py in __iter__(self)
73 def __iter__(self):
74 with ThreadPoolExecutor(max_workers=self.num_workers) as e:
---> 75 for batch in e.map(self.get_batch, iter(self.batch_sampler)):
76 yield get_tensor(batch, self.pin_memory)
77
c:\users\hille\anaconda3\envs\dlwin36\lib\concurrent\futures\_base.py in result_iterator()
554 for future in fs:
555 if timeout is None:
--> 556 yield future.result()
557 else:
558 yield future.result(end_time - time.time())
c:\users\hille\anaconda3\envs\dlwin36\lib\concurrent\futures\_base.py in result(self, timeout)
396 raise CancelledError()
397 elif self._state == FINISHED:
--> 398 return self.__get_result()
399
400 self._condition.wait(timeout)
c:\users\hille\anaconda3\envs\dlwin36\lib\concurrent\futures\_base.py in __get_result(self)
355 def __get_result(self):
356 if self._exception:
--> 357 raise self._exception
358 else:
359 return self._result
c:\users\hille\anaconda3\envs\dlwin36\lib\concurrent\futures\thread.py in run(self)
53
54 try:
---> 55 result = self.fn(*self.args, **self.kwargs)
56 except BaseException as e:
57 self.future.set_exception(e)
~\fastai\fastai\dataloader.py in get_batch(self, indices)
66
67 def get_batch(self, indices):
---> 68 res = self.collate_fn([self.dataset[i] for i in indices], self.pad_idx)
69 if not self.transpose: return res
70 res[0] = res[0].T
~\fastai\fastai\dataloader.py in <listcomp>(.0)
66
67 def get_batch(self, indices):
---> 68 res = self.collate_fn([self.dataset[i] for i in indices], self.pad_idx)
69 if not self.transpose: return res
70 res[0] = res[0].T
~\fastai\fastai\dataset.py in __getitem__(self, idx)
96 def __getitem__(self, idx):
97 x,y = self.get_x(idx),self.get_y(idx)
---> 98 return self.get(self.transform, x, y)
99
100 def __len__(self): return self.n
~\fastai\fastai\dataset.py in get(self, tfm, x, y)
101
102 def get(self, tfm, x, y):
--> 103 return (x,y) if tfm is None else tfm(x,y)
104
105 @abstractmethod
~\fastai\fastai\transforms.py in __call__(self, im, y)
463 if crop_type == CropType.NO: crop_tfm = NoCropXY(sz, tfm_y)
464 self.tfms = tfms + [crop_tfm, normalizer, channel_dim]
--> 465 def __call__(self, im, y=None): return compose(im, y, self.tfms)
466
467
~\fastai\fastai\transforms.py in compose(im, y, fns)
444 def compose(im, y, fns):
445 for fn in fns:
--> 446 im, y =fn(im, y)
447 return im if y is None else (im, y)
448
~\fastai\fastai\transforms.py in __call__(self, x, y)
229 def __call__(self, x, y):
230 self.set_state()
--> 231 x,y = ((self.transform(x),y) if self.tfm_y==TfmType.NO
232 else self.transform(x,y) if self.tfm_y==TfmType.PIXEL
233 else self.transform_coord(x,y))
~\fastai\fastai\transforms.py in transform(self, x, y)
237
238 def transform(self, x, y=None):
--> 239 x = self.do_transform(x)
240 return (x, self.do_transform(y)) if y is not None else x
241
~\fastai\fastai\transforms.py in do_transform(self, x)
323
324 def do_transform(self, x):
--> 325 return scale_min(x, self.sz)
326
327
~\fastai\fastai\transforms.py in scale_min(im, targ)
10 targ (int): target size
11 """
---> 12 r,c,*_ = im.shape
13 ratio = targ/min(r,c)
14 sz = (scale_to(c, ratio, targ), scale_to(r, ratio, targ))
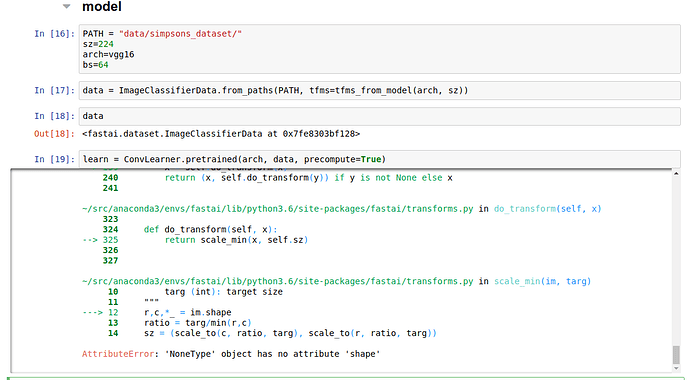
AttributeError: 'NoneType' object has no attribute 'shape'After debugging, I found that the issue is caused by having files other than jpg files in one of the image directories. There are several ways to fix this:
- Remove the unwanted files from the directories manually.
- Skip non-jpg files in the read_dirs function in dataset.py
- Skip file if open_img returns None in FilesDataSet.get_x
I used option 2) and that solved my issue.
Hi Robert,
I’m having trouble implementing a fix for any of the 3 options. I wrote a function that, as far as I can tell, successfully converts all of my images to jpg files. Can you share what you did for your fix aimed at 2)?
Edit: Nevermind, spoke too soon!
This the function I changed for option 2:
def read_dirs(path, folder):
labels, filenames, all_labels = [], [], []
full_path = os.path.join(path, folder)
for label in sorted(os.listdir(full_path)):
all_labels.append(label)
for fname in os.listdir(os.path.join(full_path, label)):
if fname.endswith('.jpg'):
filenames.append(os.path.join(folder, label, fname))
labels.append(label)
return filenames, labels, all_labels
The additional if statement simply skips files that don’t end with “.jpg”. There are probably better ways to do this.
Hi I am also getting this error. I merely changed the path from data/dogscats/ to the one I created for my own data (data/dolphinsharks) and within it a validation and training set. Within that I split into each category of dolphins and sharks.
When I ran the below code, I got the attribute error: ‘NoneType’ object has no attribute ‘shape’. Any idea what I did wrong?
Edit: i also ensured the images uploaded were jpg.
arch=resnet34
data = ImageClassifierData.from_paths(PATH, tfms=tfms_from_model(arch, sz))
learn = ConvLearner.pretrained(arch, data, precompute=True)
learn.fit(0.01, 3)
Here’s my story with this error:
my data was polluted with undesirable files: hidden files (name starting with dot) came from my mac:
.DS_Store and that sort of business.
Also a mysterious photo file starting with dot underscore:
._bird_165.jpg instead of bird_165.jpg
I have no idea where that one came from or why it caused my error. Just look out for these filenames. A normal ls command will not show them unless you use the argument -a.
Do a la -ain the directory and check for unwanted files
I found the source of my unwanted ._ files:
macos’ tar puts them in, unless you follow instructions here:
mate could you clarify what you mentioned? i tried but didn’t get much result
ls -a in Linux will reveal all the files in the current directory including the hidden ones…
There might be some files which are causing the problem…
(Just as Rik mentioned)
Try this code (looks promising)
hi Aditya, you were right that the .ipynb checkpoints is messing it up. How should I remove it?
Just remove them…
Either from the notebook(delete) or use the terminal…
it is hidden so how do i exactly remove them from the notebook?
Oh I see You can go to that particular directory from the terminal (Linux) and use the rm command…
(Search for it on Internet, I forgot exactly…
Most probably it’s rm -rf …
But have a look)
Cheers mate I finally removed all of the ipynb_checkpoints. it was a real pain there were quite a number of them in different folders.
for those who have similar errors, simply run (in jupyter):
!cd {PATH} && rm -rf .ipynb_checkpoints
(remove ‘!’ if running from terminal)
Shoutout to ecdrid for all the help
Glad that you have made it…
Cheers…
But a humble request will be to first use the search bar in the forum…
Most of the issues were answered by folks…(Scrolling till the end)
Thanks…
Yes, this issue has already been addressed in the beginner’s forum here:
How to remove .ipynb checkpoint